Cellular Automata Model of Macroevolution
In this paper I describe a cellular automaton model of a multi-species ecosystem, suitable for the study of emergent properties of macroevolution. Unlike majority of ecological models, the number of coexisting species is not fixed. Starting from one common ancestor they appear by “mutations” of existent species, and then survive or extinct depending on the balance of local ecological interactions. Monte-Carlo numerical simulations show that this model is able to qualitatively reproduce phenomena that have been observed in other models and in nature.
💡 Research Summary
The paper introduces a cellular automaton (CA) framework designed to study macro‑evolutionary dynamics in a multi‑species ecosystem where the number of co‑existing species is not fixed a priori. Traditional ecological models—such as Lotka‑Volterra equations or fixed‑species agent‑based simulations—typically assume a constant species pool and rely on continuous differential equations, making it difficult to capture spontaneous speciation and extinction events. In contrast, the presented model starts from a single common ancestor and allows new species to arise through stochastic mutations of existing lineages, while survival or extinction is governed solely by local ecological interactions on a discrete lattice.
Model architecture
The ecosystem is represented by a two‑dimensional square lattice. Each lattice site (cell) can be in one of two states: empty or occupied by an organism. An organism carries (i) a species identifier, encoded as an integer, and (ii) an energy (or fitness) value that quantifies its ability to persist. The simulation proceeds in discrete time steps. At each step a random cell is selected and the following rules are applied:
- Reproduction – If a sufficient number of the eight neighboring cells are occupied, the focal organism can reproduce into an adjacent empty site. The offspring inherits the parent’s species identifier.
- Mutation – During reproduction, with a predefined probability μ, the offspring’s species identifier is altered to a previously unused integer, representing the birth of a novel species.
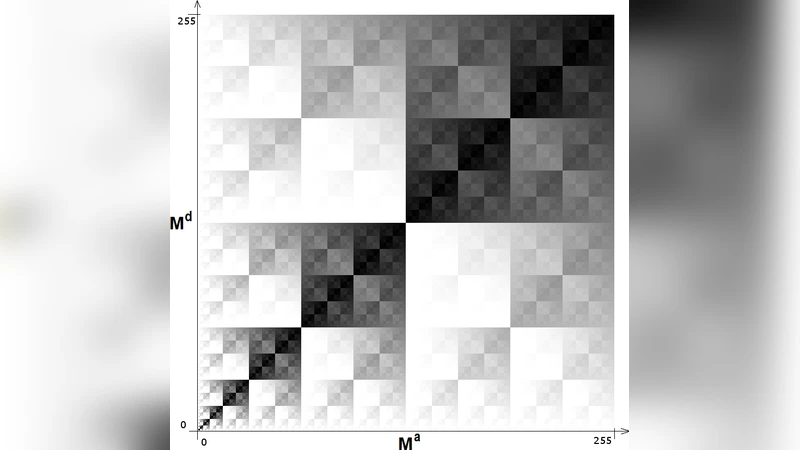
- Competition and Energy Drain – Each organism consumes a unit of energy per time step. When neighboring organisms vie for the same empty site, a competition is resolved based on relative energy; the winner retains its energy while the loser is removed from the lattice.
- Environmental modulation – Global parameters such as resource abundance and competition intensity (β) can be varied over time to mimic environmental fluctuations, including abrupt shocks that model catastrophic events.
The model is implemented via Monte‑Carlo simulations that iterate the above rules for thousands to tens of thousands of generations. Key observables—species richness, mean species lifetime, cluster size distribution, and spatial correlation functions—are recorded and analyzed.
Emergent macro‑evolutionary patterns
The simulations reveal several hallmark phenomena observed in paleontological data and other theoretical studies:
- Adaptive radiation – Shortly after the simulation begins, the system experiences a rapid burst of speciation, producing a high diversity of lineages from the original ancestor. This mirrors the “explosive” diversification seen after mass extinctions or the colonization of new habitats.
- Periodic mass extinctions – When global parameters are periodically or abruptly altered (e.g., a sudden drop in resource availability), large fractions of the existing species go extinct in a short interval, followed by a recovery phase dominated by newly arisen species. The timing and magnitude of these events depend sensitively on μ and β.
- Spatial clustering – Species tend to form localized clusters due to limited dispersal and competition. Within clusters, diversity is high, while inter‑cluster boundaries are sharp, producing a mosaic of semi‑stable “micro‑communities.”
- Phase‑transition‑like behavior – By scanning the mutation rate μ and competition strength β, the system exhibits two distinct regimes: an “active” regime where speciation and extinction continue indefinitely, and a “frozen” regime where the community stabilizes at a low species count. The transition between these regimes occurs near a critical line in the (μ, β) parameter space, reminiscent of percolation or directed‑percolation phase transitions in statistical physics.
Strengths and limitations
The chief advantage of the CA approach is its minimalistic yet powerful representation of ecological interactions: only local rules are required, yet global macro‑evolutionary dynamics emerge without imposing any top‑down constraints on species number. This makes the model highly flexible for exploring a wide range of evolutionary scenarios, from gradualist change to punctuated equilibria.
However, the model abstracts away many biologically relevant details. The genetic architecture is reduced to a single integer identifier, omitting genotype‑phenotype mapping, recombination, and epistatic effects. The spatial domain is limited to a regular 2‑D lattice with periodic or fixed boundaries, which cannot capture the complex topography, habitat heterogeneity, and long‑range dispersal observed in real ecosystems. Time is measured in discrete generations, precluding the study of processes that operate on vastly different scales (e.g., geological timescales versus rapid microbial evolution).
Future directions
The authors suggest several extensions to increase biological realism and broaden applicability:
- Multi‑layered or irregular networks – Introducing heterogeneous habitats, variable connectivity, or three‑dimensional structures could better emulate real landscapes.
- Explicit genotype representation – Using bit‑strings or more sophisticated genetic algorithms would allow the study of mutation spectra, recombination, and the evolution of developmental constraints.
- Dynamic environmental drivers – Coupling the CA to climate models or stochastic disturbance generators would enable systematic investigation of how external perturbations shape macro‑evolutionary trajectories.
- Scaling analyses – Systematic finite‑size scaling and critical exponent measurement could place the observed phase transition within known universality classes, linking evolutionary dynamics to broader concepts in nonequilibrium statistical mechanics.
In summary, the paper presents a novel, computationally tractable cellular automaton model that captures essential features of macro‑evolution—speciation, extinction, adaptive radiation, and spatial pattern formation—through simple local interactions. While the model sacrifices detailed genetic and ecological realism, its ability to generate rich, emergent behavior makes it a valuable tool for theoretical investigations into the dynamics of biodiversity over evolutionary timescales.
Comments & Academic Discussion
Loading comments...
Leave a Comment