Systems approaches and algorithms for discovery of combinatorial therapies
Effective therapy of complex diseases requires control of highly non-linear complex networks that remain incompletely characterized. In particular, drug intervention can be seen as control of signaling in cellular networks. Identification of control parameters presents an extreme challenge due to the combinatorial explosion of control possibilities in combination therapy and to the incomplete knowledge of the systems biology of cells. In this review paper we describe the main current and proposed approaches to the design of combinatorial therapies, including the empirical methods used now by clinicians and alternative approaches suggested recently by several authors. New approaches for designing combinations arising from systems biology are described. We discuss in special detail the design of algorithms that identify optimal control parameters in cellular networks based on a quantitative characterization of control landscapes, maximizing utilization of incomplete knowledge of the state and structure of intracellular networks. The use of new technology for high-throughput measurements is key to these new approaches to combination therapy and essential for the characterization of control landscapes and implementation of the algorithms. Combinatorial optimization in medical therapy is also compared with the combinatorial optimization of engineering and materials science and similarities and differences are delineated.
💡 Research Summary
The paper reviews contemporary and emerging strategies for designing combinatorial drug therapies aimed at controlling the highly non‑linear signaling networks that underlie complex diseases such as cancer, neurodegeneration, and infectious disorders. It begins by highlighting the fundamental difficulty of the problem: the space of possible drug combinations grows exponentially with the number of agents, while our knowledge of intracellular network topology and dynamics remains incomplete. Traditional clinical practice relies on empirical methods—synergy screens, dose‑equivalence adjustments, and sequential therapy trials—that are limited by small‑scale experiments, lack of mechanistic insight, and an inability to account for patient‑specific network variability.
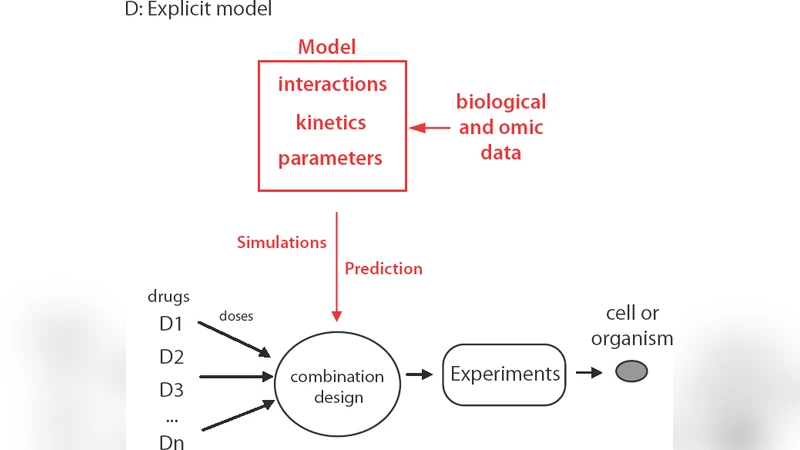
To overcome these constraints, the authors discuss systems‑biology‑driven approaches that first construct quantitative network models (ordinary differential equations, Boolean logic, Bayesian networks) using curated pathway databases and high‑throughput experimental data. Recognizing that such models are inevitably uncertain, the review emphasizes techniques for uncertainty quantification, Bayesian parameter inference, and robust optimization that explicitly incorporate model ambiguity into the design process.
A central conceptual contribution is the notion of a “control landscape.” In this framework, each point in a high‑dimensional space corresponds to a specific drug combination and dosage vector (the control parameters), while the height of the landscape represents a cost function measuring distance from a desired cellular state (e.g., apoptosis, inhibition of proliferation). Mapping this landscape enables identification of global minima (optimal therapeutic regimens) and local minima (sub‑optimal but potentially clinically viable regimens).
Algorithmic strategies for navigating the control landscape are surveyed in detail. Sparse grid sampling and Bayesian optimization are presented as efficient ways to explore the combinatorial space without exhaustive enumeration. Network‑centric heuristics—such as selecting nodes with high betweenness or degree centrality—reduce the candidate set by focusing on “control hubs.” Hybrid approaches combine dynamic simulations of the network with virtual screening to predict the effect of untested combinations. Robust optimization methods further ensure that the selected regimen performs well across a distribution of model parameters, thereby addressing inter‑patient variability.
High‑throughput technologies are portrayed as indispensable for constructing empirical control landscapes. Single‑cell RNA‑seq, phospho‑proteomics, and CRISPR‑based loss‑of‑function screens provide quantitative readouts of cellular states after drug perturbations, enabling iterative model calibration and closed‑loop optimization. This closed‑loop paradigm—alternating between computational prediction and experimental validation—offers a dramatic reduction in the number of required experiments compared with traditional open‑loop clinical trials.
The review also draws parallels with combinatorial optimization in engineering and materials science, where objectives such as alloy strength or catalyst activity are optimized over large compositional spaces. While the mathematical tools (meta‑heuristics, surrogate modeling, design of experiments) are transferable, medical applications must contend with additional layers of complexity, including biological heterogeneity, ethical constraints, and regulatory requirements.
In concluding remarks, the authors outline current bottlenecks—partial network knowledge, lack of standardized high‑throughput data pipelines, and translational hurdles—and propose a roadmap that includes the development of integrated modeling‑experimental platforms, community‑wide data standards, and multidisciplinary collaborations among biologists, clinicians, engineers, and computer scientists. By leveraging quantitative control theory, robust optimization, and modern high‑throughput measurement, the field can move toward rationally designed, patient‑specific combination therapies that are both effective and scalable.
Comments & Academic Discussion
Loading comments...
Leave a Comment