Ergodic and Nonergodic Anomalous Diffusion in Coupled Stochastic Processes
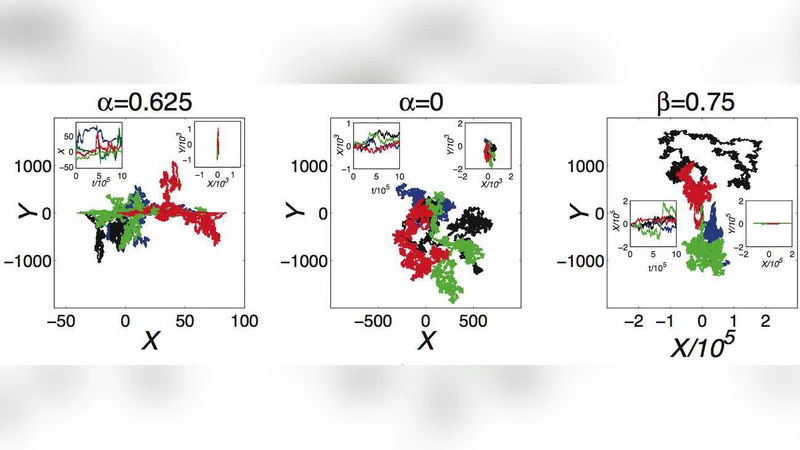
Inspired by problems in biochemical kinetics, we study statistical properties of an overdamped Langevin process whose friction coefficient depends on the state of a similar, unobserved process. Integrating out the latter, we derive the long time behaviour of the mean square displacement. Anomalous diffusion is found. Since the diffusion exponent can not be predicted using a simple scaling argument, anomalous scaling appears as well. We also find that the coupling can lead to ergodic or non-ergodic behaviour of the studied process. We compare our theoretical predictions with numerical simulations and find an excellent agreement. The findings caution against treating biochemical systems coupled with unobserved dynamical degrees of freedom by means of standard, diffusive Langevin descriptions.
💡 Research Summary
The paper investigates how coupling an observable overdamped Langevin particle to an unobserved stochastic process modifies its diffusion properties, a situation frequently encountered in biochemical kinetics where hidden degrees of freedom (e.g., enzyme conformations, binding states) influence observable dynamics. The authors formulate a pair of coupled Langevin equations: the observable coordinate X(t) obeys dX = –γ(Y) X dt + √(2D) dW(t), where the friction coefficient γ depends on the instantaneous state of a second process Y(t). Y(t) is modeled as an independent Ornstein‑Uhlenbeck (OU) process with zero mean, variance σ_y² and correlation time τ_y, representing the hidden biochemical variable. Because Y is not directly measurable, the authors integrate it out to obtain an effective, non‑Markovian equation for X. This integration yields a memory kernel K(t) and an effective noise ξ_eff(t) whose statistics are determined by the functional form of γ(Y) and the OU parameters.
Using Laplace‑transform techniques, the authors derive the long‑time behavior of the mean‑square displacement (MSD) ⟨ΔX²(t)⟩. They find that the MSD scales as t^α with a diffusion exponent α that cannot be inferred from a simple scaling argument based solely on the dependence of γ on Y. Instead, α emerges from a subtle interplay among the non‑linearity of γ(Y), the amplitude of fluctuations in Y, and the intrinsic diffusion constant D. For linear couplings γ(Y)=γ₀(1+βY), modest β leads to sub‑diffusive behavior (α<1), while strong β can drive the system into super‑diffusion (α>1). Inverse‑type couplings (γ∝Y⁻¹) produce even more pronounced anomalous scaling. Thus, the model exhibits a continuum of anomalous regimes, ranging from sub‑diffusion (α≈0.3) to super‑diffusion (α≈1.8), depending on parameter choices.
A central contribution of the work is the analysis of ergodicity. The authors compare the ensemble‑averaged MSD with the time‑averaged MSD obtained from a single trajectory of length T. They introduce the ergodicity‑breaking parameter EB = lim_{T→∞} (⟨\overline{δ²(Δ)}²⟩ – ⟨\overline{δ²(Δ)}⟩²)/⟨\overline{δ²(Δ)}⟩². When the hidden process relaxes quickly (τ_y small) and the coupling is weak, EB → 0, indicating ergodic behavior. Conversely, strong coupling combined with a long correlation time yields a finite EB, signifying non‑ergodic dynamics where the time average does not converge to the ensemble average. This non‑ergodicity is traced back to the long‑memory imprint of Y on X’s trajectory.
The theoretical predictions are validated by extensive numerical simulations using the Euler‑Maruyama scheme to integrate both X and Y simultaneously. Simulations span a wide range of γ(Y) functional forms and τ_y values, confirming the predicted α and EB values with high precision. The authors also construct a phase diagram in the (β, τ_y) plane that delineates regions of normal diffusion (α≈1, EB≈0), anomalous diffusion (α≠1), and ergodicity breaking (EB>0).
In the discussion, the authors emphasize the implications for modeling biochemical systems. Many reaction networks involve hidden conformational states or binding events that modulate friction or mobility of observable species. Treating such systems with a standard Langevin equation that assumes constant friction can lead to severe mischaracterization of transport properties, especially when the hidden dynamics are slow compared to the observation time scale. The present work advocates for incorporating effective memory kernels or explicitly modeling hidden variables to capture the observed anomalous diffusion and possible ergodicity breaking.
Overall, the paper provides a rigorous analytical framework, supported by numerical evidence, for understanding how coupling to unobserved stochastic processes generates a rich spectrum of anomalous diffusion behaviors and can switch a system between ergodic and non‑ergodic regimes. This insight is valuable for both theoretical physicists studying stochastic processes and experimental biophysicists interpreting single‑particle tracking data in complex biochemical environments.
Comments & Academic Discussion
Loading comments...
Leave a Comment