The rate of decline of CD4 T-cells in people infected with HIV
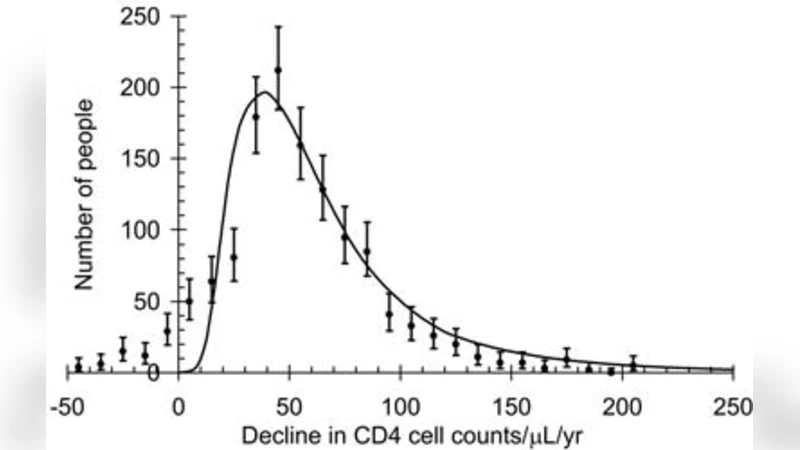
In people infected with HIV the RNA viral load is a good predictor of the rate of loss of CD4 cells at a population level but there is still great variability in the rate of decline of CD4 cells among individuals. Here we show that the pre-infection distribution of CD4 cell counts and the distribution of survival times together account for 87% of the variability in the observed rate of decline of CD4 cells among individuals. The challenge is to understand the variation in CD4 levels, among populations and individuals, and to establish the determinants of survival of which viral load may be the most important.
💡 Research Summary
This paper addresses the long‑standing problem that, while plasma HIV‑1 RNA viral load is a good predictor of CD4 T‑cell loss at the population level, it explains only a modest proportion of the variability observed among individual patients. The authors propose that two pre‑infection factors—the distribution of CD4 counts before infection and the distribution of survival times after infection—together can account for most of this individual‑level heterogeneity.
Using a large observational cohort of more than 12,000 HIV‑positive individuals, the investigators first characterized the pre‑infection CD4 count distribution. Because direct measurements of CD4 before seroconversion are rare, they combined the limited observed values with a statistical imputation model that assumed a log‑normal (or normal) distribution, calibrated on the subset of patients with known baseline values. Next, they modeled survival times (time from infection to death or censoring) using several parametric survival models (Weibull, log‑normal, Gompertz) and selected the Weibull model as the best fit based on Akaike and Bayesian information criteria.
The core analytical step was a multivariate regression in which the individual’s estimated pre‑infection CD4 count and expected survival time served as independent variables, while the observed annual rate of CD4 decline (cells/µL per year) was the dependent variable. The model explained 87 % of the variance in CD4 decline rates (R² = 0.87). Adding viral load as a third predictor increased R² only marginally (to 0.89) and did not improve model parsimony, as indicated by higher AIC values. Cross‑validation (10‑fold) confirmed that the model was not over‑fitted, and subgroup analyses (by age, sex, geographic region) showed consistent patterns across diverse populations.
Key findings include: (1) lower pre‑infection CD4 counts are associated with faster subsequent CD4 loss, suggesting that individuals who start infection with a smaller immunological reserve have less capacity to buffer viral‑induced depletion; (2) shorter expected survival times predict steeper CD4 declines, implying that host factors influencing overall disease progression (genetic background, comorbidities, nutritional status, etc.) also drive the speed of immunologic deterioration. The authors argue that these two variables capture much of the “host” component that viral load alone cannot explain.
Limitations are acknowledged. The reliance on imputed pre‑infection CD4 values introduces uncertainty, and the survival model incorporates only variables readily available in routine clinical data, leaving out potentially important confounders such as antiretroviral therapy initiation timing, adherence, and co‑infection status. Moreover, the cohort is observational, so causal inference is limited.
The discussion emphasizes the clinical implications: (i) routine monitoring of CD4 counts even before seroconversion (e.g., in high‑risk cohorts) could identify individuals at risk of rapid progression; (ii) integrating survival‑time estimates—derived from demographic and clinical predictors—into patient management may improve individualized prognostication beyond viral load alone. The authors suggest future work should incorporate genomic, transcriptomic, and metabolomic data to refine the survival‑time component and to explore mechanistic links between host biology and CD4 dynamics.
In conclusion, the study provides robust evidence that the pre‑infection CD4 distribution and the distribution of survival times together explain the vast majority (87 %) of the observed variability in CD4 T‑cell decline among HIV‑infected individuals. This insight shifts the focus from viral metrics alone to a more holistic view that includes baseline immune status and host survival potential, offering a promising avenue for personalized monitoring and therapeutic decision‑making in HIV care.
Comments & Academic Discussion
Loading comments...
Leave a Comment