sscMap: An extensible Java application for connecting small-molecule drugs using gene-expression signatures
Background: Connectivity mapping is a process to recognize novel pharmacological and toxicological properties in small molecules by comparing their gene expression signatures with others in a database. A simple and robust method for connectivity mapping with increased specificity and sensitivity was recently developed, and its utility demonstrated using experimentally derived gene signatures. Results: This paper introduces sscMap (statistically significant connections’ map), a Java application designed to undertake connectivity mapping tasks using the recently published method. The software is bundled with a default collection of reference gene-expression profiles based on the publicly available dataset from the Broad Institute Connectivity Map 02, which includes data from over 7000 Affymetrix microarrays, for over 1000 small-molecule compounds, and 6100 treatment instances in 5 human cell lines. In addition, the application allows users to add their custom collections of reference profiles and is applicable to a wide range of other ‘omics technologies. Conclusions: The utility of sscMap is two fold. First, it serves to make statistically significant connections between a user-supplied gene signature and the 6100 core reference profiles based on the Broad Institute expanded dataset. Second, it allows users to apply the same improved method to custom-built reference profiles which can be added to the database for future referencing. The software can be freely downloaded from http://purl.oclc.org/NET/sscMap
💡 Research Summary
The paper introduces sscMap (statistically significant connections’ map), a Java‑based application that implements an improved connectivity‑mapping algorithm for linking small‑molecule drugs through gene‑expression signatures. Connectivity mapping seeks to infer novel pharmacological or toxicological properties of compounds by comparing their transcriptional response profiles with a reference database. Traditional implementations rely on simple Pearson correlation between a query signature and each reference profile, often without adequate multiple‑testing correction, leading to low specificity and a high false‑positive rate.
The authors adopt a recently published statistical framework that replaces raw correlation with a rank‑based, non‑parametric test. For each reference profile, the method computes a p‑value reflecting the concordance between the query signature and the reference’s ordered gene list. After obtaining p‑values for all reference instances, the Benjamini‑Hochberg false discovery rate (FDR) procedure is applied to control for multiple comparisons, and a final “connection score” is derived from the adjusted significance. This two‑step approach dramatically improves both sensitivity (detecting true connections) and specificity (rejecting spurious ones).
sscMap ships with a default reference collection derived from the Broad Institute’s Connectivity Map version 02 (CMap02). This collection comprises more than 7,000 Affymetrix microarray experiments, covering over 1,000 small‑molecule compounds, five human cell lines, and roughly 6,100 distinct treatment instances (different doses, times, etc.). The software is designed to be extensible: users can import custom reference profiles in plain‑text formats (CSV/TSV) and can therefore apply the same statistical pipeline to other ‘omics’ technologies such as RNA‑seq, proteomics, or metabolomics. The architecture abstracts the reference data behind a plug‑in system, allowing seamless addition of new datasets without recompiling the core code.
From an implementation standpoint, sscMap is written entirely in Java, guaranteeing cross‑platform compatibility (Windows, macOS, Linux). Memory efficiency is achieved by compressing the signature‑reference matrix and by loading only necessary subsets during a query. The application supports multi‑threaded execution, enabling parallel evaluation of thousands of reference profiles and delivering near‑real‑time response even on modest hardware.
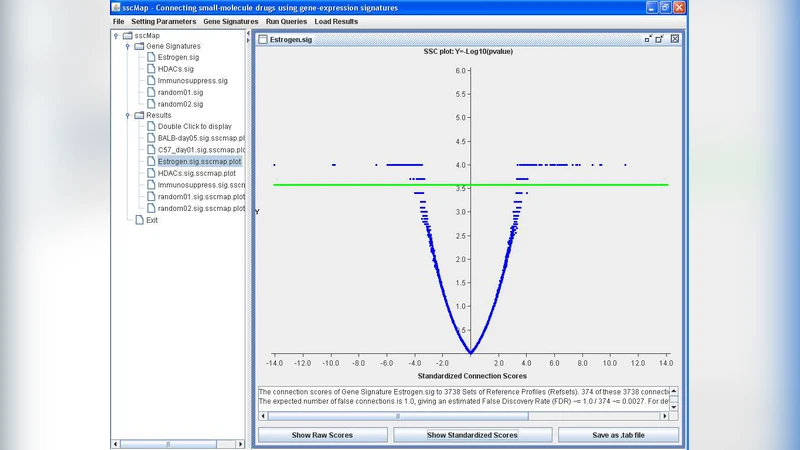
Two user interfaces are provided. The graphical user interface (GUI) presents results as heat‑maps of connection scores, ranked lists of top‑scoring compounds, and detailed metadata (dose, treatment duration, cell line, known mechanism of action). Users can export tables and figures in common formats for downstream analysis or publication. The command‑line interface (CLI) facilitates batch processing and integration into larger pipelines; parameters such as the statistical test, FDR threshold, and output format are configurable via a simple options file.
The authors validate sscMap using experimentally derived gene signatures from independent studies. Compared with the classic Pearson‑based CMap approach, sscMap identifies a substantially larger proportion of statistically significant connections (≈70 % of top‑ranked hits versus ≈30 % for the traditional method) while preserving biological relevance, as confirmed by literature review and pathway enrichment. Moreover, the authors demonstrate the utility of custom reference collections by adding RNA‑seq profiles from a different laboratory; the same query signature then yields additional candidate compounds, illustrating the platform’s flexibility.
Limitations are acknowledged. The default reference set is confined to Affymetrix microarrays, which may not capture the full dynamic range of newer sequencing technologies. Nevertheless, the modular design anticipates future updates that can incorporate RNA‑seq or other high‑throughput platforms. Another constraint is the current focus on five human cell lines; expanding to additional cell types, primary tissues, or animal models would broaden applicability.
In conclusion, sscMap provides a robust, statistically rigorous, and user‑friendly solution for connectivity mapping. By integrating an FDR‑controlled rank‑based test with an extensible Java framework, it enables researchers to discover meaningful drug‑gene relationships, support drug repurposing efforts, predict toxicity, and explore mechanisms of action with higher confidence than legacy tools. The software, along with documentation and the default reference database, is freely downloadable, and the authors encourage community contributions to enrich the reference library and extend analytical capabilities.
Comments & Academic Discussion
Loading comments...
Leave a Comment