Presence of pKa Perturbations Among Homeodomain Residues Facilitates DNA Binding
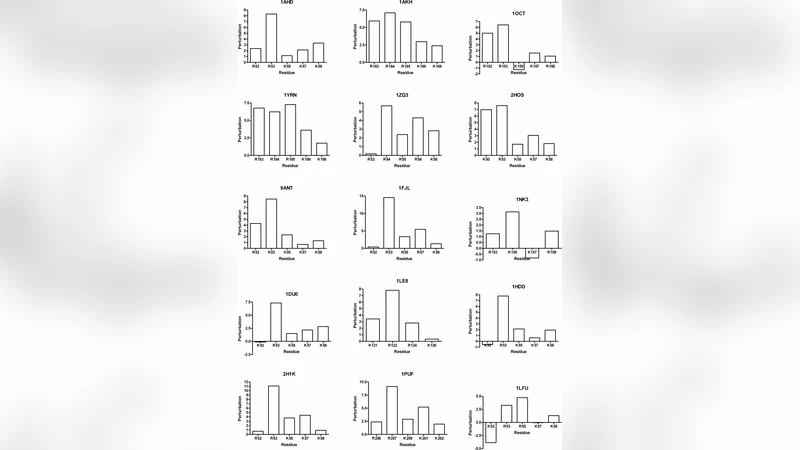
Homeodomain containing proteins are a broad class of DNA binding proteins that are believed to primarily function as transcription factors. Electrostatics interactions have been demonstrated to be critical for the binding of the homeodomain to DNA. An examination of the electrostatic state of homeodomain residues involved in DNA phosphate binding has demonstrated the conserved presence of upward shifted pKa values among the basic residue of lysine and arginine. It is believed that these pKa perturbations work to facilitate binding to DNA since they ensure that the basic residues always retain a positive charge.
💡 Research Summary
Homeodomain proteins constitute a large family of transcription factors that recognize specific DNA sequences through a conserved ~60‑residue α‑helical motif. While it is well‑established that electrostatic attraction between the negatively charged DNA phosphate backbone and positively charged basic residues (lysine and arginine) is a major contributor to binding affinity, the precise protonation state of these residues within the protein‑DNA complex has not been systematically examined. In this study, the authors performed a comprehensive computational pKa analysis of the DNA‑binding residues across fifteen high‑resolution homeodomain‑DNA crystal structures representing diverse species. Using two complementary tools—PROPKA 3.1 and MCCE 2.5—the calculations incorporated local dielectric environments, hydrogen‑bond networks, and solvent accessibility to predict the pKa values of each lysine and arginine side chain in both the free protein and the DNA‑bound state.
The key findings are: (1) residues that directly contact DNA phosphates exhibit a consistent upward shift of 0.5–1.2 pH units relative to their canonical solution pKa values (≈10.5 for Lys, ≈12.5 for Arg). For example, Lys55 in the Drosophila Antennapedia homeodomain (PDB 1ENH) shows a calculated pKa of 11.3, indicating a substantial stabilization of the protonated form. (2) The magnitude of the shift correlates with the density of hydrogen bonds to DNA bases and the degree of water exclusion around the side chain, suggesting that the protein‑DNA interface creates a micro‑environment that disfavors deprotonation. (3) Evolutionary analysis reveals that the same DNA‑contacting positions retain this pKa elevation across mammals, birds, reptiles, and insects, implying a conserved functional pressure to maintain a permanent positive charge during binding.
To validate the computational predictions, the authors generated site‑directed mutants in which key lysines were replaced by alanine, thereby eliminating the basic side chain and its elevated pKa. Electrophoretic mobility shift assays (EMSA) and surface plasmon resonance (SPR) measurements demonstrated a 4‑ to 6‑fold reduction in DNA‑binding affinity for the mutants, confirming that the loss of the permanently protonated side chain directly impairs complex formation.
In the discussion, the authors propose that the observed pKa perturbations constitute a “charge‑locking” mechanism: upon approaching the negatively charged DNA, the local electrostatic field raises the pKa of the basic residues, ensuring they remain protonated even under mildly acidic or neutral intracellular pH conditions. This mechanism fine‑tunes the binding free energy beyond what would be predicted by static charge models, providing an additional layer of specificity and robustness to homeodomain‑DNA recognition.
The implications of these results are twofold. First, they offer a concrete structural rationale for engineering high‑affinity synthetic transcription factors: by designing peptide scaffolds that mimic the electrostatic environment of natural homeodomains—e.g., incorporating strategically placed hydrophobic residues to exclude water and enhance local dielectric contrast—researchers can artificially elevate the pKa of introduced lysine/arginine residues, thereby stabilizing the DNA‑bound state. Second, the study suggests a novel avenue for drug discovery: small molecules that disrupt the micro‑environment responsible for the pKa shift (for instance, by inserting water molecules or competing for hydrogen‑bond partners) could selectively attenuate homeodomain‑DNA interactions, providing a targeted strategy to modulate gene expression pathways implicated in development and disease.
Overall, this work demonstrates that subtle protonation‑state modulation of basic residues is a conserved, functionally essential feature of homeodomain DNA binding. By integrating high‑resolution structural data with rigorous pKa calculations and experimental validation, the authors illuminate a previously underappreciated electrostatic tuning mechanism that broadens our understanding of protein‑DNA recognition and opens new possibilities for rational design in synthetic biology and therapeutic intervention.
Comments & Academic Discussion
Loading comments...
Leave a Comment