Phyllotaxis, a model
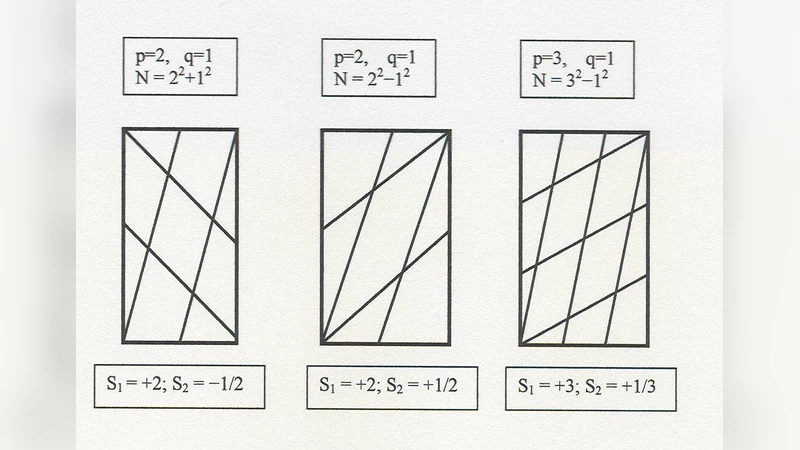
A model of the regular arrangement of leaves on a plant stem (phyllotactic patterns) is proposed, based on a new plant pattern algorithm. Tripartite patterning is proposed to occur by the interaction of two signaling pathways. Each pathway produces stimulated extracellular emission of like ligand upon activation of its respective receptor, as well as inhibiting such emission from the other pathway. Patterns arise spontaneously from zero density of activated receptor. All known phyllotactic patterns are reproduced, Fibonacci, distichous, decussate, and whorls, as well as the rare monostichy, with only one leaf directly above the previous.
💡 Research Summary
The paper introduces a novel, biologically grounded model for phyllotaxis—the regular arrangement of leaves on a plant stem—by invoking the interaction of two mutually inhibitory signaling pathways. Each pathway comprises a specific cell‑surface receptor (designated R_A and R_B) and its cognate extracellular ligand (L_A and L_B). When a receptor is activated, it triggers the production and secretion of its own ligand, establishing a positive feedback loop that amplifies the pathway’s activity. Simultaneously, the ligand of one pathway suppresses the ligand production of the opposite pathway, creating a cross‑inhibitory circuit.
Mathematically, the system is described by a set of coupled reaction‑diffusion equations. The densities of activated receptors, a(x,t) and b(x,t), evolve according to activation functions f_a(l_A) and f_b(l_B) and inhibition functions g_a(l_B) and g_b(l_A), while the ligand concentrations l_A and l_B follow production‑degradation dynamics with rates α and β and diffuse with coefficient D_l. The equations are:
∂a/∂t = f_a(l_A) – g_a(l_B) + D_a∇²a
∂b/∂t = f_b(l_B) – g_b(l_A) + D_b∇²b
∂l_A/∂t = α_a a – β_a l_A + D_l∇²l_A
∂l_B/∂t = α_b b – β_b l_B + D_l∇²l_B
The model starts from a completely quiescent state (a = b = l_A = l_B = 0) with only infinitesimal random perturbations. Linear stability analysis reveals that, for a broad region of parameter space, the uniform state becomes unstable and spatial modes with specific wave numbers grow, leading to self‑organized patterns without any pre‑imposed template.
Numerical simulations demonstrate that by modestly adjusting production rates, diffusion coefficients, and inhibition strengths, the model reproduces all classic phyllotactic arrangements observed in nature. When the cross‑inhibition is moderate and diffusion relatively fast, a spiral pattern emerges whose divergence angle converges to ~137.5°, matching the Fibonacci phyllotaxis seen in sunflowers and many other species. Stronger inhibition with slower diffusion yields a distichous (two‑row) pattern with 180° alternation, while a balanced set of parameters generates a decussate (four‑row) arrangement with 90° rotational symmetry. By further restricting diffusion, the system forms whorls (multiple leaves at the same node) and, intriguingly, a monostichy pattern where a single leaf appears directly above the previous one—an arrangement rarely reported in real plants but successfully captured by the model’s extreme parameter regime.
The authors argue that the two‑pathway scheme is biologically plausible, likening L_A and L_B to plant hormones such as auxin and cytokinin, whose transport and feedback mechanisms are well documented. They suggest that pathway A could dominate in the epidermal layer while pathway B operates in sub‑epidermal tissues, providing the spatial segregation necessary for the observed patterns.
Limitations are acknowledged. The model assumes only two pathways, whereas real plants integrate many hormonal and environmental cues. Ligand diffusion is treated as simple linear diffusion, ignoring cell wall heterogeneity, active transport, or anisotropic conductivity that are known to affect auxin distribution. Moreover, the identification of the actual molecular players (receptors, ligands) remains speculative; experimental validation would require genetic or pharmacological manipulation of candidate genes and observation of resulting phyllotactic changes.
In conclusion, this work offers a compelling synthesis of reaction‑diffusion theory and plant signaling biology, delivering a unified framework that can generate the full spectrum of known leaf arrangements from a single set of governing principles. It opens avenues for experimental testing, for extending the model to incorporate additional pathways, and for applying the underlying design logic to synthetic biology, tissue engineering, and biomimetic architecture.
Comments & Academic Discussion
Loading comments...
Leave a Comment