Effect of adhesion geometry and rigidity on cellular force distributions
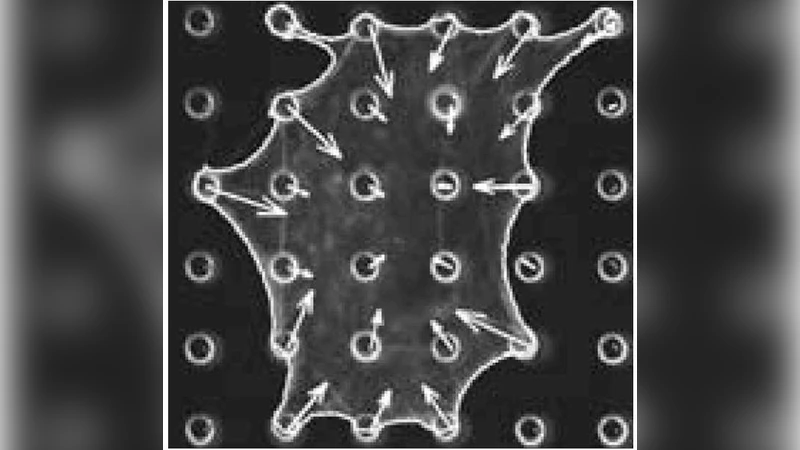
The behaviour and fate of tissue cells is controlled by the rigidity and geometry of their adhesive environment, possibly through forces localized to sites of adhesion. We introduce a mechanical model that predicts cellular force distributions for cells adhering to adhesive patterns with different geometries and rigidities. For continuous adhesion along a closed contour, forces are predicted to be localized to the corners. For discrete sites of adhesion, the model predicts the forces to be mainly determined by the lateral pull of the cell contour. With increasing distance between two neighboring sites of adhesion, the adhesion force increases because cell shape results in steeper pulling directions. Softer substrates result in smaller forces. Our predictions agree well with experimental force patterns measured on pillar assays.
💡 Research Summary
The paper investigates how the geometry of adhesive sites and the rigidity of the substrate together shape the distribution of traction forces generated by adherent tissue cells. The authors develop a mechanical model that treats the cell as a two‑dimensional elastic membrane characterized by a surface tension γ and a bending rigidity κ. Adhesion to the substrate is introduced as a boundary constraint, enforced with a Lagrange multiplier.
For continuous adhesion along a closed contour (e.g., circular or polygonal patterns), the model yields an analytical solution showing that the traction force density is not uniform but becomes highly concentrated at points where the curvature changes abruptly—namely the corners of the shape. This “corner force” prediction follows directly from minimizing the total elastic energy: the membrane can reduce bending energy by localizing deformation at discrete high‑curvature locations, which in turn requires larger reaction forces from the substrate at those points.
When adhesion is restricted to discrete spots (micropatterned pillars, adhesive islands), the cell contour between two neighboring adhesion points is approximated by a smooth spline or Bézier curve that minimizes the same energy functional. The geometry of the curve is dictated by the distance d between the adhesion sites and the angle at which the curve meets each site. As d increases, the curve must adopt a larger curvature to connect the points, which steepens the pulling direction of the membrane. The model therefore predicts a linear relationship between the magnitude of the traction force at a site and the inter‑site distance (F ∝ γ d), reflecting the increased lateral pull required to maintain the same membrane tension.
Substrate rigidity is incorporated by treating the underlying material as a linear elastic half‑space with stiffness K. The vertical displacement u of the substrate under a given traction force obeys Hooke’s law (u = F/K). Consequently, for a fixed desired deformation (set by the cell’s shape), the required traction force scales directly with K. Softer substrates (low K) thus experience smaller forces, while stiffer substrates (high K) support larger forces for the same cellular geometry.
The authors validate the model using micro‑pillar traction‑force assays. Cells are cultured on arrays of pillars arranged in various geometries (square, triangular, circular) and with different pillar stiffnesses (silicon versus polymer). High‑resolution imaging of pillar deflection provides quantitative maps of traction forces. The experimental data confirm three key predictions: (1) forces are localized at corners for continuous adhesion patterns, (2) the magnitude of forces at discrete adhesion sites grows with increasing spacing between neighboring sites, and (3) overall force magnitude decreases on softer substrates. The agreement between theory and experiment is quantitative, with model‑predicted force values falling within experimental error bars.
Beyond reproducing observed force patterns, the model offers a conceptual framework for designing adhesive environments that elicit specific mechanical cues. By selecting pattern geometry (continuous vs. discrete), controlling inter‑site distances, and tuning substrate stiffness, researchers can program the spatial distribution and magnitude of cellular traction forces. This capability is directly relevant to tissue engineering, where scaffold architecture must guide cell alignment, differentiation, and extracellular matrix deposition, as well as to disease models such as cancer invasion, where altered adhesion geometry and matrix stiffness drive aberrant force generation.
In summary, the study establishes three fundamental principles governing cellular force distribution: (i) adhesion geometry determines whether forces are corner‑localized (continuous contours) or spread according to lateral pull (discrete points); (ii) the distance between discrete adhesion sites modulates the magnitude of the forces through curvature‑dependent pulling; and (iii) substrate rigidity scales the overall force amplitude. The presented mechanical model, supported by robust experimental validation, provides a versatile tool for predicting and engineering cell‑substrate mechanical interactions across a wide range of biomedical applications.
Comments & Academic Discussion
Loading comments...
Leave a Comment