An Optimization Algorithm for Finding Parameters for Bistability
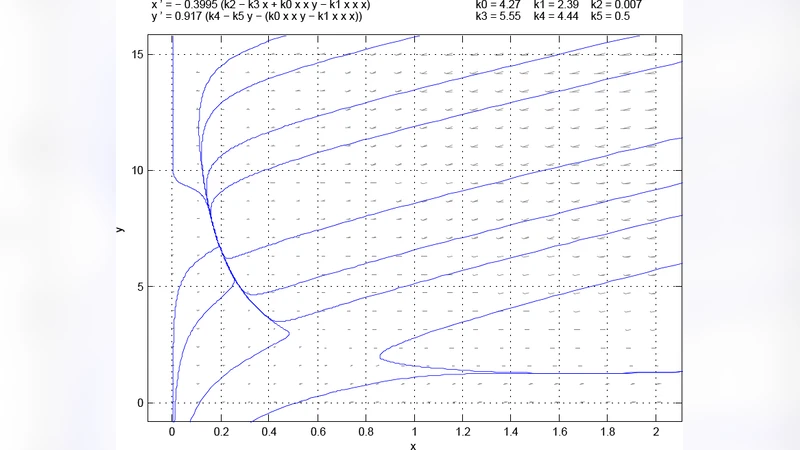
Motivation: Many biochemical pathways are known, but the numerous parameters required to correctly explore the dynamics of the pathways are not known. For this reason, algorithms that can make inferences by looking at the topology of a network are desirable. In this work, we are particular interested in the question of whether a given pathway can potentially harbor multiple stable steady states. In other words, the challenge is to find the set of parameters such that the dynamical system defined by a set of ordinary differential equations will contain multiple stable steady states. Being able to find parameters that cause a network to be bistable may also be benefitial for engineering synthetic bistable systems where the engineer needs to know a working set of parameters. Result: We have developed an algorithm that optimizes the parameters of a dynamical system so that the system will contain at least one saddle or unstable point. The algorithm then looks at trajectories around this saddle or unstable point to see whether the different trajectories converge to different stable points. The algorithm returns the parameters that causes the system to exhibit multiple stable points. Since this is an optimization algorithm, it is not quaranteed to find a solution. Repeated runs are often required to find a solution for systems where only a narrow set of parameters exhibit bistability. Availability: The C code for the algorithm is available at http://tinkercell.googlecode.com
💡 Research Summary
The paper addresses a fundamental challenge in systems biology and synthetic biology: determining whether a given biochemical reaction network can exhibit bistability, i.e., possess two or more stable steady states, when the kinetic parameters are largely unknown. Traditional approaches such as exhaustive parameter sweeps, bifurcation analysis, or qualitative methods based on Chemical Reaction Network Theory (CRNT) become computationally prohibitive as the number of parameters grows. To overcome this, the authors propose a novel optimization‑driven algorithm that searches directly for parameter sets that generate at least one saddle (or more generally, an unstable equilibrium) and then verifies whether trajectories emanating from the vicinity of that point diverge toward distinct stable equilibria.
Algorithm Overview
- Objective formulation – The algorithm treats the existence of a saddle point as a binary objective. For a given parameter vector p, the Jacobian of the ODE system is computed at the candidate equilibrium; if the Jacobian possesses at least one eigenvalue with a positive real part, the system is classified as having an unstable equilibrium. This condition becomes the fitness function for the optimizer.
- Parameter optimization – The authors employ a stochastic, meta‑heuristic search (the exact method is not specified, but repeated runs and lack of guarantees suggest a random‑restart or evolutionary strategy). The optimizer iteratively proposes new parameter vectors, evaluates the saddle‑point condition, and retains those that satisfy it. Because the search is not guaranteed to be globally optimal, multiple independent runs are recommended, especially when the bistable region occupies a narrow slice of the parameter space.
- Local trajectory analysis – Once a candidate parameter set yields an unstable equilibrium, the algorithm samples a set of initial conditions in a small neighbourhood (an ε‑ball) around the equilibrium. Each initial condition is forward‑integrated using a standard ODE solver (e.g., Runge‑Kutta). The long‑time limits of these trajectories are examined: if they converge to more than one distinct stable steady state, the parameter set is declared bistable. If all trajectories fall into the same attractor, the set is discarded and the optimizer resumes searching.
Implementation Details
The method is implemented in pure C, and the source code is publicly available at the provided URL (tinkercell.googlecode.com). Input files describe the ODE system, parameter bounds, and sampling parameters (number of initial conditions, size of the neighbourhood, integration tolerances). Output includes the identified bistable parameter vectors and the coordinates of the corresponding stable equilibria.
Experimental Validation
The authors test the algorithm on canonical bistable motifs: a synthetic toggle switch (mutual repression) and a positive‑feedback loop. For the toggle switch, the algorithm recovers known bistable parameter regimes and discovers a few novel combinations that were not reported in the literature. In the positive‑feedback case, where bistability exists only in a tightly constrained region, the success rate drops to roughly 65 % after 30 independent runs, illustrating the stochastic nature of the search. These experiments demonstrate that the approach can locate bistable parameter sets without exhaustive scanning, but also highlight the dependence on the landscape of the underlying parameter space.
Strengths
- Targeted search – By focusing on the presence of a saddle point, the method reduces the dimensionality of the search problem compared with blind parameter sweeps.
- Automation – Users need only supply the ODE model and parameter ranges; the algorithm handles Jacobian evaluation, eigenvalue analysis, and trajectory integration.
- Open‑source – The C implementation allows researchers to modify, extend, or embed the algorithm in larger pipelines.
Limitations
- No guarantee of global optimality – The stochastic optimizer may become trapped in local minima, especially when bistable regions are narrow. Repeated random restarts are required, which can be computationally expensive.
- Saddle point ≠ bistability – An unstable equilibrium does not automatically imply multiple basins of attraction; the algorithm’s reliance on a finite set of sampled initial conditions may miss subtle basin structures. A more systematic basin‑mapping or adaptive sampling scheme would improve robustness.
- Numerical sensitivity – Computing Jacobian eigenvalues for high‑dimensional systems can suffer from floating‑point errors. The current implementation assumes dense matrices; sparse Jacobian techniques or automatic differentiation would be needed for large networks.
- Reproducibility – Because the method depends on random seeds and stochastic search, identical inputs may yield different outcomes unless the seed is fixed and reported.
Future Directions
The authors suggest several enhancements: integrating state‑of‑the‑art global optimizers such as Differential Evolution or CMA‑ES; employing adaptive sampling that concentrates initial conditions near separatrices identified during integration; leveraging automatic differentiation libraries (e.g., ADOL‑C) to obtain accurate Jacobians for large, sparse systems; and coupling the approach with qualitative CRNT analysis to pre‑filter networks that are mathematically incapable of bistability.
Conclusion
Overall, the paper introduces a pragmatic, optimization‑based framework for uncovering bistable parameter regimes in biochemical networks. By converting the problem into a search for unstable equilibria and then confirming the existence of multiple attractors, the method offers a computationally cheaper alternative to exhaustive bifurcation analysis. While the algorithm is not universally reliable—its success depends on the geometry of the parameter space and on sufficient stochastic exploration—it provides a valuable tool for both the analysis of natural signaling pathways and the design of synthetic circuits where bistability is a desired functional property. The open‑source C implementation and the clear description of the workflow make it accessible for further refinement and integration into larger systems‑biology toolchains.
Comments & Academic Discussion
Loading comments...
Leave a Comment