Optimal flux states, reaction replaceability and response to knockouts in the human red blood cell
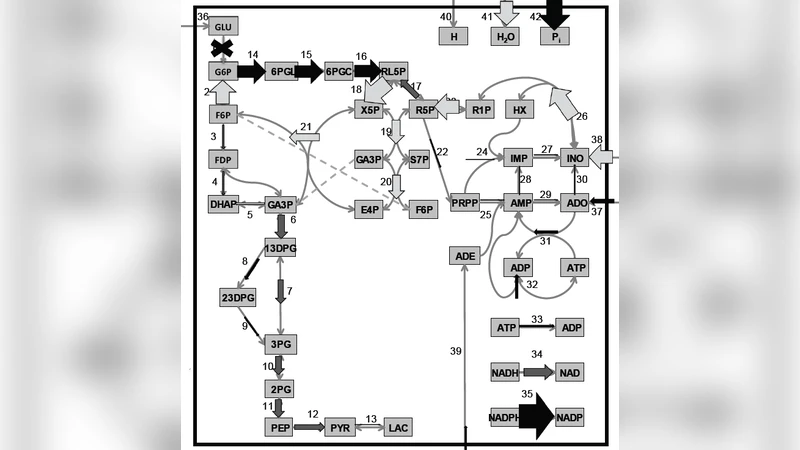
Characterizing the capabilities, criticalities and response to perturbations of genome-scale metabolic networks is a basic problem with important applications. A key question concerns the identification of the potentially most harmful knockouts. The integration of combinatorial methods with sampling techniques to explore the space of viable flux states may provide crucial insights on this issue. We assess the replaceability of every metabolic conversion in the human red blood cell by enumerating the alternative paths from substrate to product, obtaining a complete map of the potential damage of single enzymopathies. Sampling the space of optimal flux states in the healthy and in the mutated cell reveals both correlations and complementarity between topologic and dynamical aspects.
💡 Research Summary
The paper tackles two intertwined problems in the systems‑level analysis of the human red blood cell (RBC) metabolic network: (i) how to quantify the replaceability of each biochemical conversion, i.e., the extent to which a reaction can be bypassed by alternative routes, and (ii) how the space of optimal flux distributions differs between a healthy cell and cells carrying single‑enzyme knockouts. To address (i), the authors start from a curated genome‑scale RBC model comprising roughly 50 reactions and 30 internal metabolites. For every reaction they enumerate all simple, non‑cyclic paths that connect the substrate to the product, imposing a maximum path length of ten steps to keep the combinatorial explosion tractable. Each discovered path contributes to a “replaceability score,” essentially the weighted count of alternative routes. This metric goes beyond traditional binary essentiality analyses by capturing the richness of possible metabolic rerouting.
For (ii), the authors formulate a series of linear programming (LP) problems that reflect physiologically relevant objectives, such as maximal ATP yield or optimal oxygen‑delivery efficiency. Because the LP solutions are not unique, the authors employ a Hit‑and‑Run (HR) Monte‑Carlo sampler to draw uniformly from the convex polytope of all flux vectors that satisfy the stoichiometric, thermodynamic, and capacity constraints while also achieving the chosen optimum. The resulting ensemble of flux states provides a probability distribution for each reaction’s flux, enabling a direct comparison between flux variability (dynamic flexibility) and replaceability (topological flexibility).
The study proceeds to compare the “wild‑type” RBC with several single‑gene knockouts that model common enzymopathies: glucose‑6‑phosphate dehydrogenase (G6PD) deficiency, hexokinase (HK) loss, and pyruvate kinase (PK) deficiency. In the wild‑type, ATP production is robust, NADPH generation is balanced, and the flux distributions are relatively broad, reflecting the network’s intrinsic redundancy. In the G6PD‑deficient model, the replaceability score for the G6PD reaction is low, and the sampled fluxes show a tightly constrained NADPH flux that collapses, forcing the cell to activate low‑efficiency side pathways (e.g., the 2,3‑BPG shunt) to maintain redox balance. Conversely, HK and PK reactions possess higher replaceability scores because multiple alternative routes (e.g., via phosphoglucomutase or the pentose phosphate pathway) can compensate for their loss; accordingly, their fluxes exhibit larger variability across the sampled optimal states. Notably, even when ATP output remains near‑optimal in PK‑deficient simulations, the redistribution of fluxes reveals a shift toward upstream glycolytic steps, highlighting a subtle metabolic re‑wiring that would be invisible to a simple flux‑balance analysis without sampling.
A key contribution of the work is the construction of a “risk matrix” that integrates replaceability and flux variability. Reactions are classified into high‑risk (low replaceability, low flux variability), medium‑risk, and low‑risk categories. High‑risk reactions—such as G6PD, transaldolase, and certain transport steps—are predicted to be the most damaging when knocked out, aligning with clinical observations of severe hemolytic anemia in G6PD deficiency. Medium‑risk reactions include those that can be bypassed but only through routes that impose a metabolic cost, while low‑risk reactions are both topologically and dynamically flexible. The matrix provides a systematic way to prioritize candidate drug targets or metabolic engineering interventions: for instance, augmenting NADPH regeneration pathways could mitigate G6PD deficiency, whereas stimulating alternative glycolytic branches might alleviate PK deficiency.
Overall, the study demonstrates that a combined topological‑dynamic framework yields a richer, more predictive picture of metabolic robustness than either approach alone. By enumerating alternative pathways and sampling the optimal flux polytope, the authors uncover hidden compensatory mechanisms, quantify the severity of potential enzymopathies, and propose a practical tool for risk assessment in metabolic disease and synthetic biology. The methodology is readily extensible to other cell types and larger genome‑scale models, offering a valuable addition to the toolbox of systems biologists and computational biochemists.
Comments & Academic Discussion
Loading comments...
Leave a Comment