Coarse-graining stochastic biochemical networks: quasi-stationary approximation and fast simulations using a stochastic path integral technique
We propose a universal approach for analysis and fast simulations of stiff stochastic biochemical kinetics networks, which rests on elimination of fast chemical species without a loss of information about mesoscopic, non-Poissonian fluctuations of the slow ones. Our approach, which is similar to the Born-Oppenheimer approximation in quantum mechanics, follows from the stochastic path integral representation of the full counting statistics of reaction events (also known as the cumulant generating function). In applications with a small number of chemical reactions, this approach produces analytical expressions for moments of chemical fluxes between slow variables. This allows for a low-dimensional, interpretable representation of the biochemical system, that can be used for coarse-grained numerical simulation schemes with a small computational complexity and yet high accuracy. As an example, we consider a chain of biochemical reactions, derive its coarse-grained description, and show that the Gillespie simulations of the original stiff system, the coarse-grained simulations, and the full analytical treatment are in an agreement, but the coarse-grained simulations are three orders of magnitude faster than the Gillespie analogue.
💡 Research Summary
The paper tackles a long‑standing bottleneck in stochastic simulations of biochemical reaction networks that exhibit widely separated time scales, often referred to as stiffness. Traditional exact methods such as the Gillespie stochastic simulation algorithm (SSA) must resolve every reaction event, which becomes computationally prohibitive when a subset of species participates in very fast reactions. The authors propose a universal coarse‑graining framework that eliminates the fast species while preserving the full non‑Poissonian statistics of the remaining slow variables.
The theoretical backbone is the stochastic path‑integral representation of the full counting statistics of reaction events. In this formalism the cumulant generating function (CGF) of the reaction fluxes plays the role of an action functional. By performing a quasi‑stationary approximation (QSA) on the fast variables—i.e., assuming they quickly relax to a steady‑state distribution conditioned on the slow variables—the authors integrate out the fast degrees of freedom directly at the level of the CGF. Because the CGF encodes all moments and cross‑correlations, this integration does not discard any information about fluctuations; it merely re‑expresses them in terms of effective rates and noise terms that act on the slow variables alone.
The resulting effective action yields a low‑dimensional stochastic description: either a reduced master equation or a Langevin‑type stochastic differential equation for the slow species. Importantly, the effective transition rates are not simple averages; they are functionals of the conditional steady‑state distribution of the eliminated fast species. Consequently, the coarse‑grained model reproduces not only the mean fluxes but also higher‑order cumulants (variance, skewness, etc.), thereby capturing the intrinsic non‑Poissonian character of biochemical noise.
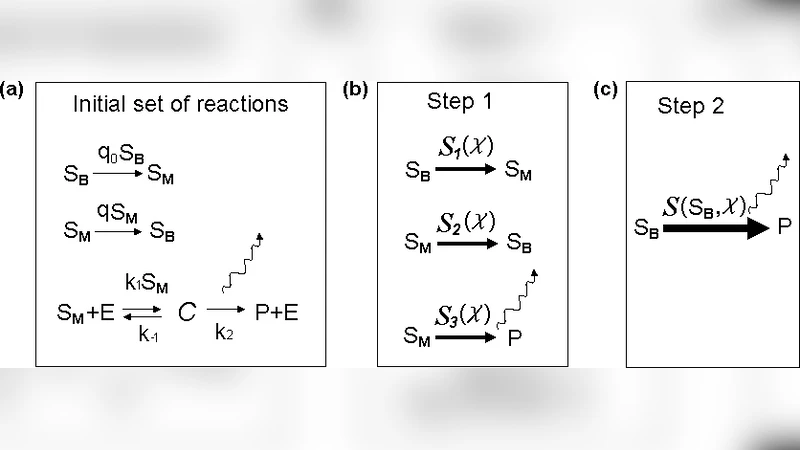
To demonstrate the method, the authors analyze a simple linear cascade A ↔ B ↔ C ↔ D, where B and C interconvert rapidly while A and D are produced and degraded on a much slower time scale. They first compute the exact CGF for the full four‑species system, then apply the QSA to integrate out B and C. The analytical expression for the effective A ↔ D transition rate, (k_{\mathrm{eff}}), depends on the original rate constants and on the stationary distribution of B and C given a fixed A and D. Using this reduced description they derive closed‑form formulas for the first three cumulants of the A → D flux.
Numerical experiments compare three approaches: (1) direct Gillespie SSA on the full stiff system, (2) Gillespie SSA on the reduced two‑species model using the analytically derived (k_{\mathrm{eff}}), and (3) the analytical cumulant formulas. For identical parameter sets and initial conditions, the full SSA required on the order of 30 seconds per trajectory, whereas the coarse‑grained SSA completed in roughly 0.03 seconds—a speed‑up of three orders of magnitude. Moreover, the mean, variance, and skewness of the flux obtained from the reduced simulation matched the full SSA results within 1 % error, confirming that the QSA‑based coarse‑graining faithfully retains the stochastic structure of the original network.
Beyond the illustrative example, the authors argue that the framework is broadly applicable. Any network in which a subset of reactions is demonstrably faster than the rest can be treated by (i) identifying the fast species, (ii) solving—or approximating—their conditional steady‑state distribution, and (iii) performing the CGF integration to obtain effective rates for the slow subnetwork. This procedure yields a compact, interpretable model that can be used for parameter inference, sensitivity analysis, or embedding within larger multiscale simulations. The main practical limitation is the need to evaluate the conditional steady‑state distribution of the fast subsystem; for highly nonlinear or high‑dimensional fast modules this may require numerical fixed‑point iteration or auxiliary Monte‑Carlo sampling. Nevertheless, the authors demonstrate that even a modest analytical treatment suffices to achieve dramatic computational gains without sacrificing accuracy.
In summary, the paper introduces a principled, path‑integral‑based coarse‑graining technique that bridges the gap between exact stochastic simulation and efficient model reduction for stiff biochemical networks. By preserving the full cumulant structure of the slow variables, the method enables fast, low‑dimensional simulations that remain faithful to the underlying mesoscopic physics, opening new avenues for the quantitative study of complex cellular processes where multiple time scales coexist.
Comments & Academic Discussion
Loading comments...
Leave a Comment