Principal Fitted Components for Dimension Reduction in Regression
We provide a remedy for two concerns that have dogged the use of principal components in regression: (i) principal components are computed from the predictors alone and do not make apparent use of the response, and (ii) principal components are not invariant or equivariant under full rank linear transformation of the predictors. The development begins with principal fitted components [Cook, R. D. (2007). Fisher lecture: Dimension reduction in regression (with discussion). Statist. Sci. 22 1–26] and uses normal models for the inverse regression of the predictors on the response to gain reductive information for the forward regression of interest. This approach includes methodology for testing hypotheses about the number of components and about conditional independencies among the predictors.
💡 Research Summary
The paper addresses two long‑standing criticisms of principal component regression (PCR): (i) PCR derives components solely from the predictor matrix X, ignoring any information from the response Y, and (ii) PCR is not invariant under full‑rank linear transformations of X, which can lead to different fitted models after a change of basis. To overcome these drawbacks, the authors build on the concept of Principal Fitted Components (PFC), originally introduced by Cook (2007), and develop a systematic framework that uses an inverse‑regression model of the form
X | Y ~ N( μ(Y), Σ ),
where μ(Y) = Γ f(Y). Here f(Y) is a sufficient statistic for Y (often a low‑dimensional vector of basis functions such as polynomials or splines), and Γ ∈ ℝ^{p×d} is a matrix whose column space defines a d‑dimensional reduction subspace that captures all the regression information about Y. By modeling the conditional mean of X given Y, the method explicitly incorporates the response into the construction of the components, ensuring that the resulting directions are “response‑oriented” rather than purely variance‑oriented.
Estimation proceeds via maximum likelihood. The log‑likelihood depends on Γ and the covariance matrix Σ, which is structured to separate the variation explained by the fitted part Γ f(Y) from the residual variation. Maximizing the likelihood yields estimates of Γ that are, under the normality assumption, sufficient dimension reduction (SDR) directions: any function of X that is sufficient for Y can be expressed as a function of ΓᵀX. Consequently, the fitted components are invariant to any full‑rank linear transformation A of the predictors: if X* = A X, the estimated reduction matrix becomes A⁻¹Γ, preserving the subspace and the predictive performance.
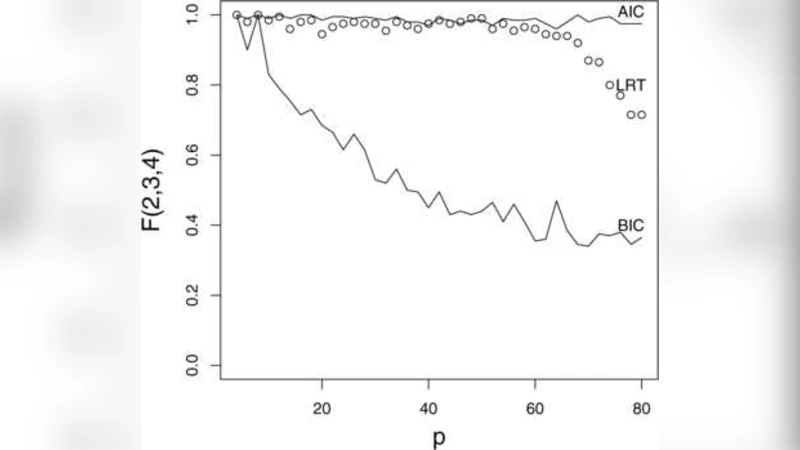
A major contribution of the paper is a formal testing procedure for the dimension d. The authors derive a likelihood‑ratio test (LRT) comparing models with d versus d − 1 components. Under the null hypothesis, the LRT statistic follows a chi‑square distribution with degrees of freedom equal to the difference in parameter counts, allowing a data‑driven choice of d. In addition, the paper provides Wald‑type and LRT procedures for testing conditional independencies among the predictors, such as X_i ⟂ X_j | Y. These tests exploit the block‑diagonal structure of Σ implied by the inverse model and give practitioners tools for model simplification and variable selection.
The methodology is illustrated through extensive simulation studies and two real‑data applications. Simulations vary the signal‑to‑noise ratio, the degree of nonlinearity between X and Y, and the dimensionality (including p ≫ n scenarios). Across all settings, PFC consistently yields lower mean‑squared prediction error and more accurate identification of the true reduction subspace than PCR, especially when the relationship between X and Y is not captured by the leading variance directions of X. The real‑data examples involve a high‑dimensional spectroscopic dataset and a genomics data set linking gene expression to a phenotype. In both cases, the authors demonstrate that a modest number of fitted components (often d = 2 or 3) suffices to achieve predictive performance comparable to or better than full‑dimensional models, while also providing interpretable linear combinations of the original variables.
The paper concludes by emphasizing the theoretical advantages of PFC: (1) explicit incorporation of response information into the dimension‑reduction step, (2) invariance under any full‑rank linear transformation of the predictors, (3) a likelihood‑based framework that yields natural hypothesis tests for the number of components and for conditional independencies, and (4) practical robustness in high‑dimensional settings. The authors argue that Principal Fitted Components constitute a powerful, statistically principled alternative to traditional PCR for regression problems where dimension reduction and interpretability are both essential.
Comments & Academic Discussion
Loading comments...
Leave a Comment