The Statistical Analysis of fMRI Data
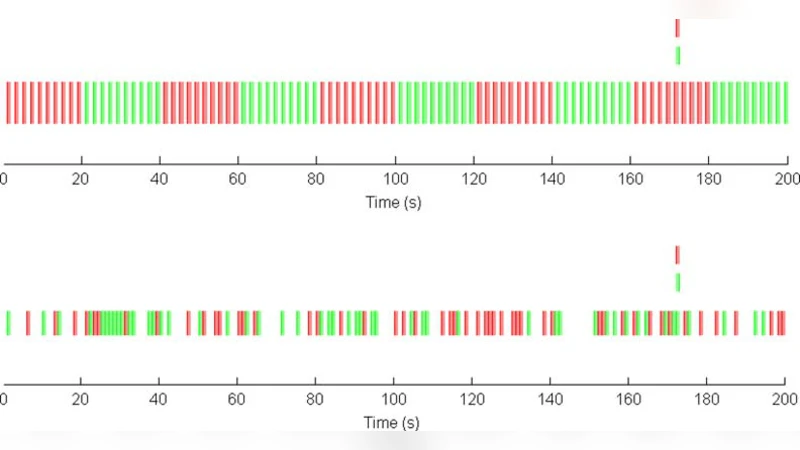
In recent years there has been explosive growth in the number of neuroimaging studies performed using functional Magnetic Resonance Imaging (fMRI). The field that has grown around the acquisition and analysis of fMRI data is intrinsically interdisciplinary in nature and involves contributions from researchers in neuroscience, psychology, physics and statistics, among others. A standard fMRI study gives rise to massive amounts of noisy data with a complicated spatio-temporal correlation structure. Statistics plays a crucial role in understanding the nature of the data and obtaining relevant results that can be used and interpreted by neuroscientists. In this paper we discuss the analysis of fMRI data, from the initial acquisition of the raw data to its use in locating brain activity, making inference about brain connectivity and predictions about psychological or disease states. Along the way, we illustrate interesting and important issues where statistics already plays a crucial role. We also seek to illustrate areas where statistics has perhaps been underutilized and will have an increased role in the future.
💡 Research Summary
The paper provides a comprehensive overview of the statistical challenges and solutions that arise throughout the entire pipeline of functional magnetic Resonance Imaging (fMRI) research, from raw signal acquisition to predictive modeling of clinical or psychological states. It begins by emphasizing that a typical fMRI experiment generates massive, high‑dimensional time series data with low signal‑to‑noise ratio and intricate spatio‑temporal correlations. Because of these properties, simple descriptive statistics are insufficient; sophisticated, interdisciplinary statistical methods are required.
In the acquisition stage, the authors discuss physical noise sources (thermal fluctuations, scanner vibrations) and physiological confounds (cardiac pulsation, respiration). They review regression‑based correction techniques such as RETROICOR and illustrate how these confounds can be modeled as separate stochastic processes (e.g., Gaussian noise for thermal effects, ARMA models for physiological cycles). To capture variability across subjects and scanning sessions, the paper advocates hierarchical linear mixed‑effects models and Bayesian multi‑level frameworks, which treat subject‑specific baselines as random effects while estimating fixed experimental effects.
Pre‑processing steps—realignment, spatial normalization, and smoothing—are examined from a statistical perspective. While smoothing enhances spatial continuity and improves the validity of Gaussian random field assumptions, it also reduces degrees of freedom and inflates the multiple‑comparison problem. The authors recommend selecting the smoothing kernel size based on the intrinsic spatial resolution of the data rather than arbitrarily, thereby balancing sensitivity and specificity.
For the core inference of brain activation, the manuscript contrasts three major approaches. The classic voxel‑wise General Linear Model (GLM) treats each voxel independently, yielding t‑statistics that are easy to interpret but ignore spatial dependence. Multivariate pattern analysis (MVPA) leverages the joint information of many voxels using classifiers such as support vector machines, offering higher detection power for distributed patterns. A spatial Bayesian model, often built on Markov Random Field priors, explicitly incorporates neighboring voxel correlations, shrinking confidence intervals and improving statistical power. The authors also discuss multiple‑comparison correction strategies, highlighting the advantages of False Discovery Rate (FDR) control and Threshold‑Free Cluster Enhancement (TFCE) over traditional Bonferroni adjustments.
Connectivity analysis is divided into functional connectivity (FC) and effective connectivity (EC). FC is typically quantified by Pearson or partial correlations, which capture static co‑activation but cannot infer directionality. To address this limitation, the paper reviews Dynamic Causal Modeling (DCM) and Structural Equation Modeling (SEM) as frameworks for estimating directed, time‑varying influences among brain regions. Bayesian Model Selection (BMS) is presented as a principled way to compare competing causal architectures and select the most plausible network. Graph‑theoretic metrics—node centrality, modularity, global efficiency—are described for summarizing the topology of the inferred networks.
Predictive modeling sections explore how machine learning and deep learning have been applied to fMRI-derived features. Traditional regularized linear models (LASSO, Elastic Net) provide interpretability and guard against overfitting, whereas nonlinear methods (support vector machines, random forests, convolutional neural networks, recurrent neural networks, graph neural networks) often achieve higher classification accuracy. The authors stress rigorous validation protocols, including k‑fold cross‑validation, bootstrap resampling, and external cohort testing, to avoid optimistic bias. Model‑explainability tools such as SHAP and LIME are recommended for translating black‑box predictions into neuroscientifically meaningful statements.
A forward‑looking segment identifies areas where statistical methodology is still under‑exploited. Real‑time fMRI neurofeedback could benefit from online Bayesian updating and reinforcement‑learning algorithms that adapt stimulation protocols on the fly. Personalized brain atlases, built via subject‑specific Bayesian priors, promise more accurate localization of functional regions for individual patients. Large‑scale open repositories (e.g., OpenNeuro, Human Connectome Project) enable meta‑analyses that require hierarchical Bayesian models to accommodate heterogeneity across studies. Finally, the paper highlights privacy‑preserving statistical techniques, such as differential privacy, as essential for sharing sensitive neuroimaging data while maintaining statistical utility.
In conclusion, the authors argue that statistics is not merely a supporting tool but a central driver of fMRI research. While much of the field still relies on voxel‑wise GLM inference, emerging Bayesian multi‑level models, dynamic causal frameworks, and integrated machine‑learning pipelines are poised to address the high dimensionality, complex dependence structures, and translational goals of modern neuroimaging. The paper calls for deeper collaboration between statisticians, neuroscientists, and computer scientists to fully harness the potential of fMRI data for both basic science and clinical applications.
Comments & Academic Discussion
Loading comments...
Leave a Comment