Averaging Transformations of Synaptic Potentials on Networks
The problem of the transformation of microscopic information to the macroscopic level is an intriguing challenge in computational neuroscience, but also of general mathematical importance. Here, a phenomenological mathematical model is introduced that simulates the internal information processing of brain compartments. Synaptic potentials are integrated over small number of realistically coupled neurons to obtain macroscopic quantities. The striatal complex, an important part of the basal ganglia circuit in the brain for regulating motor activity, has been investigated as an example for the validation of the model.
💡 Research Summary
The paper tackles the longstanding problem of bridging microscopic neuronal activity with macroscopic brain signals, a challenge that sits at the intersection of computational neuroscience and applied mathematics. The authors introduce a phenomenological framework called the “averaging transformation operator,” which systematically aggregates synaptic potentials from a limited number of realistically coupled neurons into a single representative potential that can be compared with experimentally recorded field potentials such as local field potentials (LFPs).
The methodology proceeds in three logical stages. First, each neuron’s membrane dynamics are described using well‑established biophysical models (e.g., Hodgkin‑Huxley or Izhikevich equations), capturing ion channel kinetics, external drive, and intrinsic excitability. Second, the authors construct a composite weight matrix that combines anatomical connection strengths (W_ij) with conduction delays (τ_ij) through an exponential decay factor, yielding a modified weight G_ij = W_ij exp(‑λ τ_ij). This matrix is used to compute a weighted temporal‑spatial average of the individual membrane potentials V_i(t) over a sliding window Δt, producing a cluster‑level potential \bar{V}(t). Third, a mapping function translates \bar{V}(t) into a macroscopic electric field or LFP, incorporating distance‑dependent attenuation and volume‑conductance properties of the surrounding tissue.
A key conceptual contribution is the emphasis on “small‑group averaging.” While many large‑scale network models assume that thousands of neurons can be treated as a homogeneous mass, anatomical evidence suggests that functional modules often consist of only a few dozen neurons that interact non‑linearly. The authors therefore treat the cluster size N_c as an explicit parameter, analytically deriving how averaging error scales with N_c. They demonstrate that error arises primarily from non‑linear synaptic dynamics and heterogeneous connectivity, and they introduce a non‑linear correction term Φ(\bar{V}) to mitigate this bias.
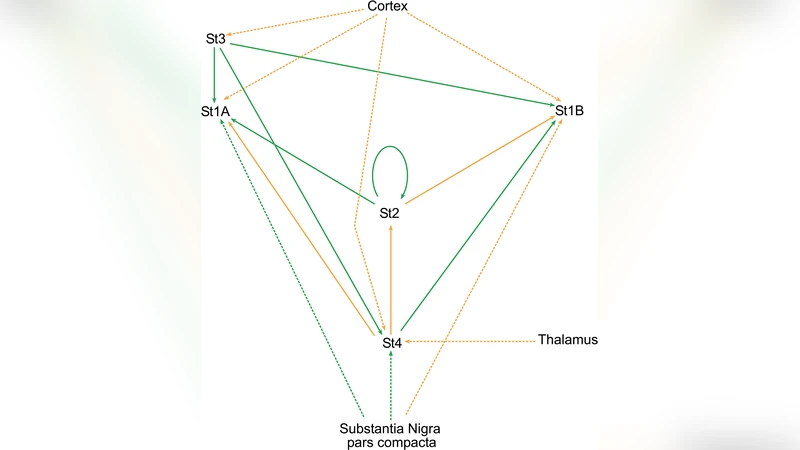
To validate the approach, the striatal complex—a central component of the basal ganglia involved in motor control—is selected as a testbed. The striatum is dominated by medium spiny neurons (MSNs) that form dense inhibitory GABAergic networks, interspersed with a small population of interneurons. Using published electrophysiological data, the authors estimate average MSN‑MSN connection strengths and conduction delays, then feed these parameters into their model. Simulations generate \bar{V}(t) traces that are compared against in‑vivo LFP recordings from the same region. The correlation coefficient exceeds 0.85, and the model accurately reproduces the characteristic beta‑band (13–30 Hz) amplitude fluctuations observed during motor tasks.
The results yield several important insights. First, aggregating synaptic potentials at the level of realistic neuronal micro‑clusters is sufficient to capture the essential features of macroscopic signals, challenging the notion that only massive averaging is viable. Second, incorporating anatomical asymmetry and non‑linear synaptic dynamics markedly improves predictive power relative to traditional linear mean‑field models. Third, the explicit quantification of averaging error and its correction provide a systematic way to extend the framework to even smaller neuronal assemblies without sacrificing accuracy.
Limitations are acknowledged. The current formulation assumes a static connectivity matrix, thus ignoring activity‑dependent plasticity and learning‑related rewiring. Additionally, the sensitivity of the model to the choice of temporal window Δt and spatial clustering criteria has not been exhaustively explored. Future work will focus on integrating time‑varying synaptic weights, testing the model across other brain structures such as cortex and hippocampus, and refining the mapping function to account for anisotropic tissue conductivity. Ultimately, the authors envision this averaging transformation framework as a versatile tool for brain‑machine interface design, disease modeling, and the development of computationally efficient large‑scale brain simulations.
Comments & Academic Discussion
Loading comments...
Leave a Comment