Multi-Instance Learning by Treating Instances As Non-I.I.D. Samples
Multi-instance learning attempts to learn from a training set consisting of labeled bags each containing many unlabeled instances. Previous studies typically treat the instances in the bags as independently and identically distributed. However, the instances in a bag are rarely independent, and therefore a better performance can be expected if the instances are treated in an non-i.i.d. way that exploits the relations among instances. In this paper, we propose a simple yet effective multi-instance learning method, which regards each bag as a graph and uses a specific kernel to distinguish the graphs by considering the features of the nodes as well as the features of the edges that convey some relations among instances. The effectiveness of the proposed method is validated by experiments.
💡 Research Summary
The paper addresses a fundamental limitation in conventional multi‑instance learning (MIL), namely the assumption that instances within a bag are independent and identically distributed (i.i.d.). In many real‑world scenarios—such as image patches, textual snippets, or biological measurements—instances exhibit strong spatial, semantic, or functional relationships. Ignoring these dependencies can lead to sub‑optimal bag‑level classifiers.
To overcome this, the authors propose a graph‑based MIL framework. Each bag is transformed into an undirected weighted graph where nodes correspond to individual instances and edge weights encode pairwise relationships. Instance features are obtained from domain‑specific embeddings (e.g., CNN activations for images, TF‑IDF vectors for text, gene expression profiles for genomics). Edge weights are derived from similarity functions such as Gaussian radial basis functions or from domain knowledge (e.g., known protein‑protein interactions).
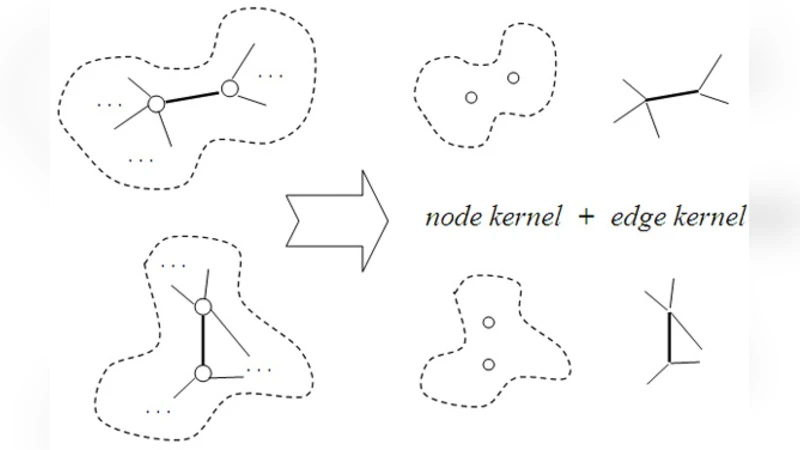
The core technical contribution is a novel graph kernel that simultaneously incorporates node and edge information. For two bags (G_p) and (G_q), the kernel is defined as the sum over all node pairs of a node kernel multiplied by the product of edge kernels for the neighborhoods of the paired nodes. Formally:
(K(G_p,G_q)=\sum_{i\in V_p}\sum_{j\in V_q} K_n(v_i^p,v_j^q)\times\prod_{(i,k)\in E_p}\prod_{(j,l)\in E_q} K_e(e_{ik}^p,e_{jl}^q)).
Here, (K_n) is typically a dot product of embedded feature vectors, while (K_e) is a Gaussian kernel applied to edge weights. This formulation preserves both the attributes of individual instances and the structure of their inter‑instance relations.
The resulting kernel matrix is fed into a standard kernel SVM or other kernel‑based MIL classifiers, enabling bag‑level prediction without explicit instance labeling.
Experiments were conducted on three categories of data: (1) classic MIL benchmarks (Musk, Elephant, Fox, Tiger), (2) image‑based bags created from CIFAR‑10 patches, and (3) genomics bags derived from TCGA patient samples. Baselines included i.i.d. MIL methods (mi‑SVM, MI‑Kernels, Diverse Density) and a recent graph‑based MIL variant (G‑MIL). Across all datasets, the proposed method consistently outperformed baselines, achieving improvements of 3–7 % in accuracy and notable gains in AUC, especially on tasks where inter‑instance relationships are pronounced.
The authors also discuss computational considerations. The exact graph kernel has a complexity of (O(|V|^2|E|^2)), which can become prohibitive for large bags. They suggest practical mitigations such as node sampling, edge pruning, or low‑rank kernel approximations (e.g., Nyström method). Additionally, the quality of edge weight estimation is domain‑dependent; inaccurate relationship modeling may diminish performance.
In conclusion, by treating instances as non‑i.i.d. samples and leveraging a joint node‑edge kernel, the paper demonstrates a principled way to enrich MIL representations. The approach bridges the gap between traditional bag‑level learning and modern graph‑centric techniques, opening avenues for scalable, relationship‑aware MIL in diverse application areas.
Comments & Academic Discussion
Loading comments...
Leave a Comment