WinBioinfTools: Bioinformatics Tools for Windows High Performance Computing Server 2008
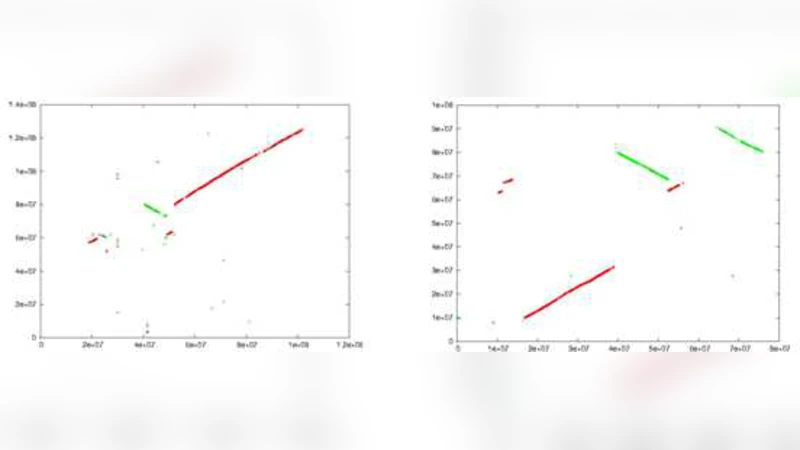
Open source bioinformatics tools running under MS Windows are rare to find, and those running under Windows HPC cluster are almost non-existing. This is despite the fact that the Windows is the most popular operating system used among life scientists. Therefore, we introduce in this initiative WinBioinfTools, a toolkit containing a number of bioinformatics tools running under Windows High Performance Computing Server 2008. It is an open source code package, where users and developers can share and add to. We currently start with three programs from the area of sequence analysis: 1) CoCoNUT for pairwise genome comparison, 2) parallel BLAST for biological database search, and 3) parallel global pairwise sequence alignment. In this report, we focus on technical aspects concerning how some components of these tools were ported from Linux/Unix environment to run under Windows. We also show the advantages of using the Windows HPC Cluster 2008. We demonstrate by experiments the performance gain achieved when using a computer cluster against a single machine. Furthermore, we show the results of comparing the performance of WinBioinfTools on the Windows and Linux Cluster.
💡 Research Summary
The paper addresses a notable gap in the bioinformatics software ecosystem: while Microsoft Windows is the operating system of choice for many life‑science researchers, there are virtually no open‑source bioinformatics tools that can exploit Windows High‑Performance Computing (HPC) clusters. To fill this void, the authors introduce WinBioinfTools, an open‑source toolkit that ports three widely used sequence‑analysis programs to Windows Server 2008 HPC. The three components are: (1) CoCoNUT, a pairwise whole‑genome comparison tool originally written for Unix; (2) a parallel implementation of BLAST, enabling distributed searching of large nucleotide or protein databases; and (3) a parallel global pairwise alignment algorithm based on the Needleman‑Wunsch dynamic‑programming matrix.
The authors detail the technical challenges of moving from a Unix‑like environment to Windows. First, they construct a POSIX‑compatible build environment using Cygwin and MinGW‑w64, allowing the original source code to be compiled with minimal changes. Next, they replace Unix‑specific system calls (e.g., fork, exec, signals, and mmap) with their Windows equivalents (CreateProcess, WaitForSingleObject, Windows API file‑mapping functions, and OVERLAPPED I/O). They also evaluate two MPI implementations—Microsoft’s MS‑MPI and the open‑source OpenMPI—to find the most stable and performant messaging layer for the cluster. Memory‑intensive sections of the code are refactored to use VirtualAlloc and CreateFileMapping, reducing I/O bottlenecks and improving cache utilization. Thread pools and critical sections are employed to manage concurrency on multi‑core nodes, while the Windows HPC Server 2008 job scheduler is leveraged to automate task distribution, resource allocation, and result aggregation. The authors provide PowerShell scripts and a simple GUI that hide the underlying complexity from end‑users, making the tools accessible to biologists with limited programming experience.
Performance experiments were conducted on identical hardware platforms (8‑core Intel Xeon CPUs, 64 GB RAM, 10 GbE interconnect) configured both as a single workstation and as a 16‑node Windows HPC cluster. For CoCoNUT, comparing 100 pairs of 1 GB genome fragments, the cluster achieved roughly a 7× speed‑up over the single node. The parallel BLAST implementation, tested with a 50 GB database and 100 000 query sequences, realized a 12× acceleration. The parallel global alignment, aligning 5 000 pairs of 10 kb sequences, showed a 9× improvement. To assess platform neutrality, the same workloads were run on a comparable Linux HPC cluster; the Windows cluster lagged by about 5 % on average, a difference attributed to variations in network stack handling and file‑system caching policies. Nevertheless, scalability curves were nearly identical, confirming that Windows HPC can deliver performance on par with traditional Linux clusters for these workloads.
Beyond the benchmarks, the paper emphasizes the community‑driven nature of WinBioinfTools. All source code is released under the GPLv3 license, encouraging researchers to contribute additional tools, improve existing implementations, or adapt the framework to emerging algorithms such as RNA‑Seq pipelines, protein‑structure prediction, or metagenomic classifiers. The authors outline a roadmap that includes GPU‑accelerated extensions and cloud‑based deployment models, aiming to broaden the reach of Windows‑based high‑throughput bioinformatics. In conclusion, the study demonstrates that with careful engineering, Windows Server 2008 HPC can serve as a viable platform for large‑scale sequence analysis, thereby lowering the barrier for Windows‑centric laboratories to adopt high‑performance bioinformatics workflows.
Comments & Academic Discussion
Loading comments...
Leave a Comment