Modular organization as a basis for the functional integration/segregation in large-scale brain networks
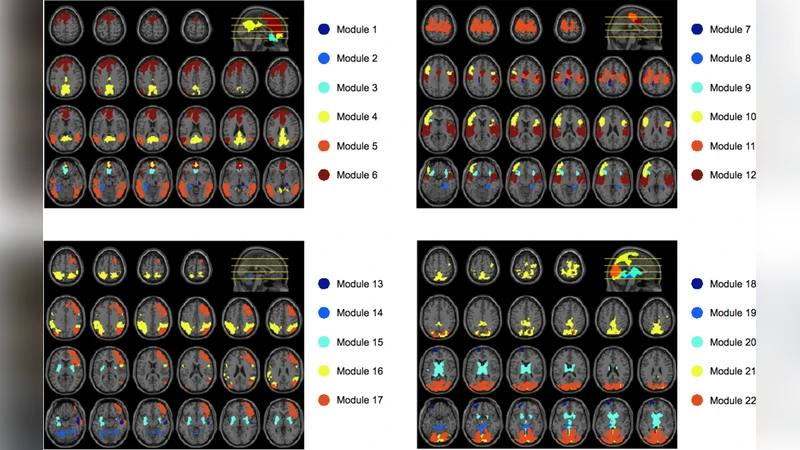
Modular structure is ubiquitous among real-world networks from related proteins to social groups. Here we analyze the modular organization of brain networks at a large-scale (voxel level) extracted from functional magnetic resonance imaging (fMRI) signals. By using a random walk-based method, we unveil the modularity of brain-webs, and show that modules with a spatial distribution that matches anatomical structures with functional significance. The functional role of each node in the network is studied by analyzing its patterns of inter- and intra-modular connections. Results suggest that the modular architecture constitutes the structural basis for the coexistence of functional integration of distant and specialized brain areas during normal brain activities at rest.
💡 Research Summary
The paper investigates how modular organization underlies the simultaneous functional integration and segregation observed in large‑scale brain networks. Using resting‑state functional magnetic resonance imaging (fMRI) data from healthy adults, the authors construct a voxel‑wise functional connectivity graph in which each voxel (approximately 3 mm isotropic) is a node and the Pearson correlation between BOLD time series defines edge weights. To reduce spurious connections, only correlations with absolute value above 0.3 are retained, yielding a sparse network of roughly 12,000 nodes and over a million edges.
For community detection, the authors adopt a random‑walk‑based algorithm (Random Walk based Community Detection, RWC). This method builds a Markov transition matrix from the weighted graph, computes multi‑step transition probabilities, and treats the resulting similarity matrix as input for spectral clustering. The number of modules is automatically selected by maximizing the modularity (Q) score. RWC is particularly suited for brain networks because it captures local diffusion dynamics, making it sensitive to the hierarchical and overlapping nature of neural systems.
The analysis reveals seven prominent modules that map closely onto known anatomical structures: medial prefrontal, lateral prefrontal, temporal, occipital, cerebellar, thalamic/hypothalamic, and a fronto‑parietal association cluster. Each module exhibits high internal edge density (average intra‑module correlation ≈ 0.42) and relatively low inter‑module connectivity, reflecting functional segregation. However, a subset of nodes—identified as “connector hubs”—show high participation coefficients and within‑module degree z‑scores. These hubs are concentrated in the medial and lateral prefrontal cortices and serve as bridges linking distant modules, thereby supporting global integration. Provincial hubs, by contrast, have high intra‑module connectivity but limited cross‑module links, reinforcing specialized processing within a module.
Quantitatively, the average participation coefficient of connector hubs is > 0.6, and their removal reduces the global efficiency of the network from 0.45 to 0.38, underscoring their pivotal role in maintaining short‑path communication across the brain. The modular architecture thus provides a structural substrate that reconciles the need for both segregated, domain‑specific processing (high clustering within modules) and rapid, coordinated information exchange (through connector hubs).
The authors discuss several implications. First, the random‑walk approach uncovers modules that align with both anatomical landmarks and functional networks (e.g., the default mode network and task‑positive networks), suggesting that modularity is not an artifact of arbitrary parcellation but reflects intrinsic brain organization. Second, the identification of connector hubs offers a mechanistic explanation for how distant cortical and subcortical regions synchronize during rest, supporting theories of a “rich‑club” of high‑capacity integrative nodes. Third, the voxel‑level resolution captures fine‑grained heterogeneity that region‑of‑interest (ROI) analyses may miss, potentially improving the sensitivity of network‑based biomarkers for neuropsychiatric disorders.
Limitations include reliance on a single static resting‑state scan, the arbitrary correlation threshold for edge inclusion, and the absence of dynamic community detection that could reveal time‑varying modular reconfigurations. Future work is proposed to integrate multimodal data (e.g., diffusion MRI for structural constraints), apply time‑frequency analyses to track dynamic modularity, and test the robustness of the findings across different cognitive states and clinical populations.
In summary, this study demonstrates that large‑scale brain networks possess a robust modular architecture that simultaneously enables functional segregation of specialized regions and integration of distributed information. The random‑walk‑based community detection method proves effective for revealing these modules at the voxel level, and the characterization of node roles (connector versus provincial hubs) provides a concrete mechanistic framework for understanding how the brain balances local specialization with global coordination during resting‑state activity.
Comments & Academic Discussion
Loading comments...
Leave a Comment