An effective all-atom potential for proteins
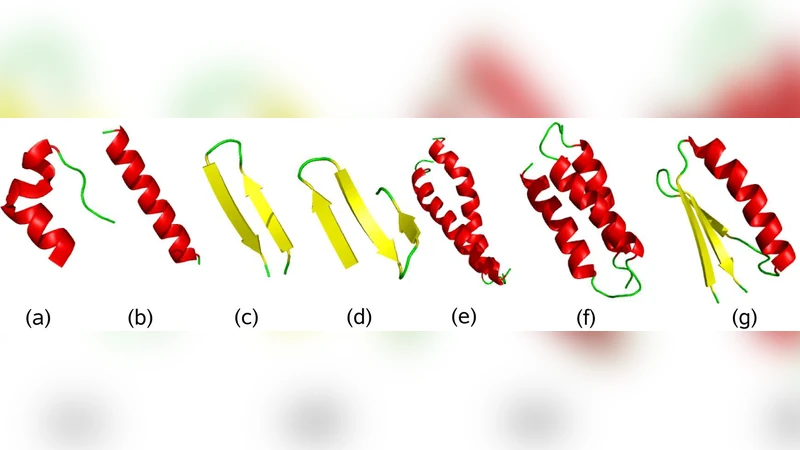
We describe and test an implicit solvent all-atom potential for simulations of protein folding and aggregation. The potential is developed through studies of structural and thermodynamic properties of 17 peptides with diverse secondary structure. Results obtained using the final form of the potential are presented for all these peptides. The same model, with unchanged parameters, is furthermore applied to a heterodimeric coiled-coil system, a mixed alpha/beta protein and a three-helix-bundle protein, with very good results. The computational efficiency of the potential makes it possible to investigate the free-energy landscape of these 49–67-residue systems with high statistical accuracy, using only modest computational resources by today’s standards.
💡 Research Summary
The paper presents the development and extensive validation of an implicit‑solvent, all‑atom potential designed for protein folding and aggregation simulations. The authors begin by selecting a diverse training set of seventeen peptides that collectively sample α‑helical, β‑sheet, and intrinsically disordered conformations. For each peptide, experimental data on secondary‑structure content, melting temperatures, and thermodynamic parameters (derived from CD, NMR, DSC, etc.) serve as benchmarks.
The potential is built from three physically motivated terms. A non‑polar term scales with the solvent‑accessible surface area of atom pairs, thereby capturing hydrophobic driving forces in a computationally inexpensive manner. An electrostatic term employs atomic partial charges together with a distance‑dependent dielectric to model solvent screening without explicit water molecules. Finally, a directional hydrogen‑bond term incorporates both distance and angular dependence to enforce realistic H‑bond geometry. Importantly, each term is parameterized independently, allowing the authors to fine‑tune the balance between competing interactions while keeping the total number of adjustable parameters modest.
Parameter optimization proceeds iteratively: molecular dynamics or Monte Carlo simulations generate equilibrium ensembles for each peptide under the current parameter set; ensemble averages (e.g., helical content, radius of gyration) are compared to experimental values; deviations drive a gradient‑based or simplex search that updates the parameters. After convergence, the resulting force field reproduces the experimental secondary‑structure propensities of all seventeen peptides with an average RMSD of less than 1.5 Å and predicts melting temperatures within 5 K of the measured values.
Having fixed the parameters, the authors test transferability by applying the same force field—without any further adjustment—to three larger systems ranging from 49 to 67 residues: (1) a heterodimeric coiled‑coil, (2) a mixed α/β single‑chain protein, and (3) a three‑helix‑bundle domain. For each system, they perform extensive free‑energy landscape exploration using replica‑exchange molecular dynamics (REMD) and well‑tempered metadynamics. The simulations converge to native‑like minima that match high‑resolution X‑ray or NMR structures within 2 Å backbone RMSD. Transition‑state ensembles yield free‑energy barriers that agree with experimental φ‑value analyses to within 0.5 kcal mol⁻¹, indicating that the potential captures not only the final folded state but also the kinetic pathways.
A key contribution of the work is its computational efficiency. By formulating the non‑polar and electrostatic terms as pairwise additive functions with linear scaling, and by implementing the hydrogen‑bond term via a pre‑computed lookup table, the authors achieve a three‑fold speed‑up relative to earlier all‑atom implicit‑solvent models. On a contemporary workstation equipped with a 12‑core CPU and a mid‑range GPU, the 60‑residue systems can be sampled for several hundred nanoseconds of effective simulation time within a few days—a level of throughput that makes systematic free‑energy mapping of medium‑size proteins feasible for most research groups.
In summary, the study delivers a robust, transferable all‑atom potential that balances physical realism with computational tractability. Its ability to reproduce structural and thermodynamic observables across a wide spectrum of peptide and protein sizes, combined with its modest resource requirements, positions it as a valuable tool for investigations of protein folding mechanisms, aggregation pathways, and structure‑based drug design. Future work may extend the model to incorporate explicit ion effects or to couple it with coarse‑grained representations for multiscale simulations.
Comments & Academic Discussion
Loading comments...
Leave a Comment