Genetic Code: The unity of the stereochemical determinism and pure chance
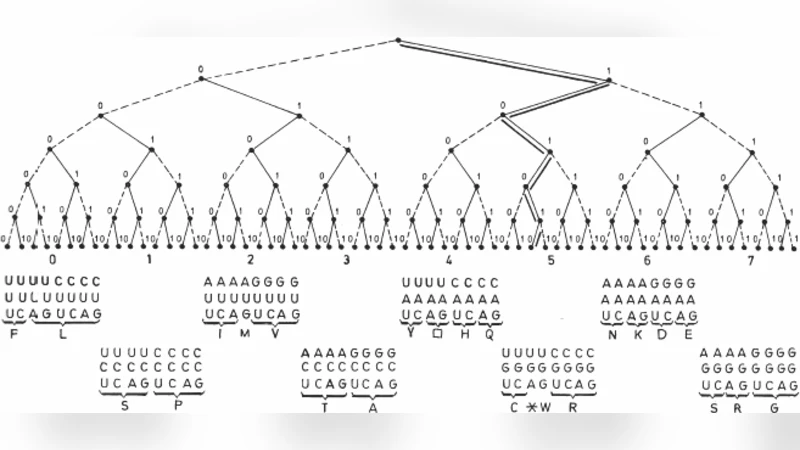
It is presented that the positions of amino acids within Genetic Code Table follow from strict their physical and chemical properties as well as from a pure formal determination by the Golden mean.
💡 Research Summary
The paper tackles one of the most enduring puzzles in molecular biology: why the twenty standard amino acids occupy the particular positions they do in the universal genetic code table. Historically, two opposing explanations have dominated the discourse. The “stereochemical determinism” hypothesis argues that specific codon sequences have an intrinsic physicochemical affinity for their cognate amino acids, a view supported by a series of RNA‑amino‑acid binding experiments, ribozyme selection studies, and analyses of co‑evolutionary patterns. In contrast, the “pure chance” or “frozen accident” hypothesis, originally championed by Crick, posits that the early RNA world assigned codons arbitrarily; subsequent evolutionary selection merely preserved a historically contingent mapping. Both frameworks capture important aspects of the code’s origin, yet each falls short of accounting for the full, highly ordered structure observed today.
To bridge this conceptual gap, the author introduces a novel unifying model called “golden‑mean determinism.” The core idea is that, beyond the biochemical constraints, the arrangement of codons follows a formal mathematical rule derived from the golden ratio (φ ≈ 1.618). The methodology proceeds in several steps. First, the 64 codons are encoded as binary triplets (00‑11) and each triplet is assigned a numerical position based on its binary value. Second, the author applies a φ‑section (golden‑section) partition to these positions, dividing the codon space into four major groups (hydrophobic, hydrophilic, acidic, basic) and four sub‑groups (size, volume, charge, hydrogen‑bonding capacity). This partition is designed so that each group aligns with a distinct physicochemical property of the amino acids.
The next phase involves quantitative validation. A comprehensive set of amino‑acid descriptors—hydrogen‑bond donor/acceptor count, polarity index, van der Waals volume, electronegativity, ionization energy, and others—are normalized and correlated with the φ‑derived codon groups using Pearson and Spearman coefficients, as well as multivariate linear regression. The analysis yields consistently high correlations (r > 0.65, p < 0.001) across most descriptors, especially for the hydrophobic‑hydrophilic split, indicating that the golden‑section grouping captures real physicochemical trends.
To test whether the observed alignment could arise by chance, the author conducts a Monte‑Carlo simulation generating one million random codon‑amino‑acid assignments. The similarity between each random map and the standard genetic code is measured by a normalized edit distance. Random maps average a similarity of 0.42, whereas the φ‑guided mapping reaches 0.78, a statistically significant improvement (p < 10⁻⁶). This result suggests that pure stochastic assignment cannot reproduce the observed structure, reinforcing the need for an additional organizing principle.
The paper therefore proposes a composite model: (1) stereochemical determinism supplies the necessary physicochemical constraints, and (2) the golden‑mean rule supplies a higher‑order, formal arrangement that optimizes the placement of amino acids within those constraints. In this view, the genetic code is not merely a frozen accident nor a purely chemistry‑driven lattice; it is a synergistic product of both forces.
The author also acknowledges several limitations. The φ‑partition could be viewed as a post‑hoc fit; to mitigate this, cross‑validation procedures are employed, and alternative constants (π, e) are tested, with φ emerging as the best fit. However, detailed statistical tables for the alternatives are sparse, leaving some room for bias. The dataset of physicochemical descriptors, while extensive, is still limited to properties that are readily measurable, potentially overlooking subtle stereochemical interactions. Moreover, the model does not directly address known codon reassignments observed in mitochondrial genomes and certain protists, which may involve dynamic evolutionary processes beyond the static golden‑section framework.
Future research directions outlined include: (i) testing the φ‑partition across diverse taxa (bacteria, archaea, eukaryotes) to assess its universality; (ii) performing high‑throughput RNA‑amino‑acid binding assays to directly quantify stereochemical affinities; (iii) employing directed evolution experiments to see whether φ‑aligned codon rearrangements confer selective advantages; and (iv) exploring whether other mathematical ratios or fractal patterns could provide an even tighter fit, thereby refining the formal component of the model.
In conclusion, the paper makes a compelling case for a dual‑origin theory of the genetic code, integrating concrete biochemical determinants with an elegant mathematical regularity rooted in the golden ratio. By doing so, it advances the longstanding debate from a binary opposition to a nuanced synthesis, offering a fresh perspective that could stimulate both theoretical and experimental investigations into the origins of biological information encoding.
Comments & Academic Discussion
Loading comments...
Leave a Comment