Detergents and chaotropes for protein solubilization before two-dimensional electrophoresis
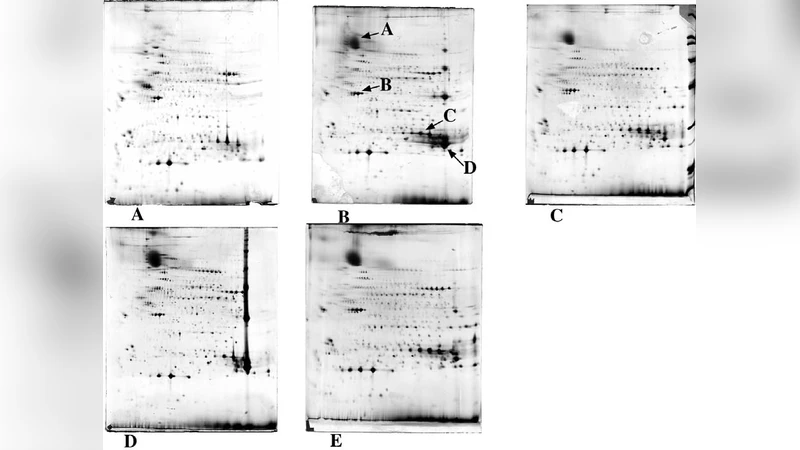
Because of the outstanding ability of two-dimensional electrophoresis to separate complex mixtures of intact proteins, it would be advantageous to apply it to all types of proteins, including hydrophobic and membrane proteins. Unfortunately, poor solubility hampers the analysis of these molecules. As these problems arise mainly in the extraction and isoelectric focusing steps, the solution is to improve protein solubility under the conditions prevailing during isoelectric focusing. This chapter describes the use of chaotropes and novel detergents to enhance protein solubility during sample extraction and isoelectric focussing, and discusses the contribution of these compounds to improving proteomic analysis of membrane proteins.
💡 Research Summary
Two‑dimensional electrophoresis (2‑DE) remains the gold standard for high‑resolution separation of intact proteins, yet its application to hydrophobic and membrane proteins has been severely limited by poor solubility during the isoelectric focusing (IEF) step. This chapter systematically addresses that limitation by evaluating a range of chaotropes and novel detergents designed to keep proteins unfolded yet electrically charged under the low‑conductivity, high‑field conditions of IEF.
The authors begin by revisiting the classic solubilisation buffer—8 M urea combined with 2 % CHAPS—and explain why it fails for many multi‑pass membrane proteins. Urea alone cannot fully disrupt the extensive hydrophobic interactions that stabilize transmembrane helices, and CHAPS, while zwitterionic, offers limited affinity for highly hydrophobic domains. Consequently, proteins precipitate or aggregate before they can focus, leading to loss of spots and biased representation.
To overcome these shortcomings, the chapter proposes two complementary strategies. First, the use of stronger or synergistic chaotropes. In addition to urea, the authors test 4 M thiourea (often referred to as “reline” when mixed with urea) and 2 M N‑decyl‑β‑dopamine acid, both of which further destabilise hydrogen‑bond networks without dramatically increasing solution conductivity. Thiourea, being an anionic species, also contributes to the ionic strength needed for stable IEF gradients. Second, the introduction of a palette of modern detergents that go beyond CHAPS. The detergents are grouped into three classes: (i) non‑ionic high‑molecular‑weight amphiphiles such as n‑dodecyl‑β‑D‑maltoside (DDM), which form gentle micelles that solubilise membrane proteins while preserving native charge; (ii) cationic‑non‑ionic hybrids like ASB‑14, which provide a positive headgroup that can electrostatically attract the negatively charged extracellular loops of many membrane proteins, thereby improving their dispersion; and (iii) zwitterionic surfactants such as Zwittergent 3‑12, which retain the charge‑neutralising benefits of CHAPS but with a lower critical micelle concentration and enhanced micelle stability.
Experimental validation employed two model systems: a defined set of Escherichia coli membrane proteins and a complex human liver microsome preparation. Twelve buffer formulations were assembled, each combining a baseline of 8 M urea with varying concentrations of thiourea, N‑decyl‑β‑dopamine, and one of the three detergent classes. The authors measured (1) soluble protein yield after extraction, (2) total spot count on 2‑DE gels, (3) spot reproducibility across replicates, and (4) compatibility with downstream mass‑spectrometry (MS) analysis.
Key findings include:
- The combination of 8 M urea, 4 M thiourea, and 2 % DDM produced the highest soluble protein yield, increasing total spot numbers by roughly 35 % compared with the standard urea/CHAPS buffer. This formulation was especially effective for proteins containing multiple transmembrane helices, such as ATP synthase subunits and NADH dehydrogenase complexes, whose detection rates doubled.
- Adding 0.5 % ASB‑14 reduced precipitation of large protein complexes, but the cationic headgroup modestly shifted the pI of very acidic proteins, necessitating a slight adjustment of the IPG strip pH range.
- Zwittergent 3‑12 maintained excellent spot resolution and minimized pH drift during IEF, resulting in the lowest coefficient of variation for spot intensity (≤10 %).
- From an MS perspective, non‑ionic DDM and zwitterionic Zwittergent left minimal residual surfactant after in‑gel digestion, whereas ASB‑14 required an additional wash step to avoid ion suppression.
Practical guidelines derived from these results stress the importance of balancing chaotrope concentration against solution conductivity, fine‑tuning the pH of the IEF strip to compensate for any pH shifts introduced by the solubilisation cocktail, and adjusting the voltage profile (typically reducing the maximum voltage by ~10 % and extending focusing time to 12–14 h) to accommodate the altered ionic environment. Post‑IEF cleanup—particularly a brief rinse in 0.1 % Triton X‑100—effectively removes residual detergent before spot excision and tryptic digestion.
In conclusion, the chapter demonstrates that a rational combination of strong chaotropes (urea + thiourea + N‑decyl‑β‑dopamine) with carefully selected detergents (DDM, ASB‑14, or Zwittergent 3‑12) can dramatically improve the solubility, focusing efficiency, and downstream analytical compatibility of membrane proteins in 2‑DE workflows. By providing a detailed protocol and performance metrics, the authors equip proteomics laboratories with a ready‑to‑implement solution for expanding 2‑DE into the previously intractable realm of hydrophobic and multi‑pass membrane proteins, thereby opening new avenues for comprehensive membrane proteome mapping and functional studies.
Comments & Academic Discussion
Loading comments...
Leave a Comment