Statistical physics of a model binary genetic switch with linear feedback
We study the statistical properties of a simple genetic regulatory network that provides heterogeneity within a population of cells. This network consists of a binary genetic switch in which stochastic flipping between the two switch states is mediated by a “flipping” enzyme. Feedback between the switch state and the flipping rate is provided by a linear feedback mechanism: the flipping enzyme is only produced in the on switch state and the switching rate depends linearly on the copy number of the enzyme. This work generalises the model of [Phys. Rev. Lett., 101, 118104] to a broader class of linear feedback systems. We present a complete analytical solution for the steady-state statistics of the number of enzyme molecules in the on and off states, for the general case where the enzyme can mediate flipping in either direction. For this general case we also solve for the flip time distribution, making a connection to first passage and persistence problems in statistical physics. We show that the statistics of the model are non-Poissonian, leading to a peak in the flip time distribution. The occurrence of such a peak is analysed as a function of the parameter space. We present a new relation between the flip time distributions measured for two relevant choices of initial condition. We also introduce a new correlation measure to show that this model can exhibit long-lived temporal correlations, thus providing a primitive form of cellular memory. Motivated by DNA replication as well as by evolutionary mechanisms involving gene duplication, we study the case of two switches in the same cell. This results in correlations between the two switches; these can either positive or negative depending on the parameter regime.
💡 Research Summary
The paper investigates a minimalist genetic regulatory circuit—a binary switch—augmented by a “flipping” enzyme whose production is contingent on the switch being in the ON state. The key novelty is the introduction of a linear feedback: the switching rate is proportional to the instantaneous copy number of the enzyme (k n), and the enzyme itself is synthesized only when the switch is ON. This framework generalizes the earlier model of PRL 101, 118104 by allowing the enzyme to catalyze flips in both directions (ON→OFF and OFF→ON) and by treating the feedback strength as a tunable parameter.
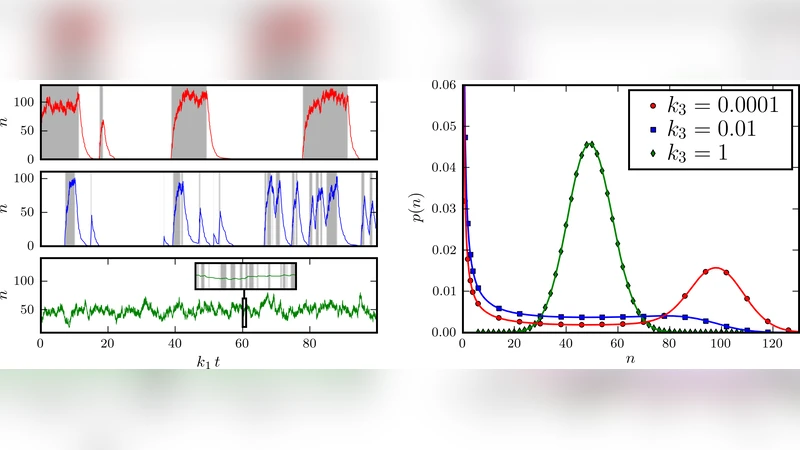
The authors formulate the stochastic dynamics through a master equation for the joint probability P(s,n,t), where s∈{0,1} denotes the switch state and n the enzyme count. By employing generating‑function techniques and Laplace transforms, they obtain closed‑form expressions for the steady‑state distributions P_ON(n) and P_OFF(n). Unlike a simple Poisson process, these distributions are non‑Poissonian: they exhibit skewed tails and a variance that grows super‑linearly with the feedback coefficient k. The mean enzyme number ⟨n⟩ and its variance are derived as explicit functions of the synthesis rate g, degradation rate γ, and feedback strength k, revealing a regime where enzyme accumulation becomes self‑reinforcing.
A central result concerns the flip‑time distribution f(τ), defined as the probability density for the interval between successive state changes. By mapping the problem onto a first‑passage time for a birth‑death process with state‑dependent rates, the authors derive an exact expression for f(τ). The distribution typically displays a pronounced peak at a characteristic time τ*≈1/(k⟨n⟩), reflecting a “waiting‑time” before enough enzyme molecules have accumulated to trigger a flip. This peak distinguishes the model from memoryless Poisson switching and provides a measurable signature of linear feedback. Moreover, the paper establishes a novel relation between flip‑time densities measured from the two possible initial conditions: f_ON(τ)=e^{-ατ} f_OFF(τ), where α is a function of the feedback parameters. This relation enables experimental cross‑validation without needing to directly observe the hidden enzyme dynamics.
Temporal correlations are quantified through the autocorrelation function C(t)=⟨σ(0)σ(t)⟩−⟨σ⟩², where σ(t) encodes the switch state. In the weak‑feedback limit C(t) decays exponentially, but as k increases the decay becomes markedly slower, producing a long‑lived tail. The authors introduce a “memory time” τ_mem, defined as the integral of C(t), and show that τ_mem scales with both the enzyme lifetime (γ⁻¹) and the feedback strength. This demonstrates that the linear feedback endows the switch with a primitive form of cellular memory, allowing a cell to retain information about its past state over many generations.
The study is further extended to a system containing two identical switches within the same cell, sharing the same pool of flipping enzyme. Even though the switches do not directly interact, the shared enzyme creates indirect coupling. Analytical treatment reveals that the inter‑switch correlation coefficient ρ can be either positive or negative depending on the parameter regime: high synthesis rates and weak feedback favor positive correlation (both switches tend to be in the same state), whereas low synthesis and strong feedback can produce negative correlation (the switches tend to be in opposite states). This dual‑switch analysis offers insight into how gene duplication and shared regulatory resources might generate diverse phenotypic outcomes in a population.
Biologically, the model captures essential features of systems such as DNA replication origins, phase‑varying promoters, and epigenetic switches where enzyme concentrations modulate transition probabilities. The non‑Poissonian statistics, peaked flip‑time distribution, and long‑range temporal correlations together provide a mechanistic explanation for observed heterogeneity and memory in bacterial persister cells, viral latency, and developmental bistability.
In conclusion, the paper delivers a complete analytical solution for a linear‑feedback binary genetic switch, elucidating how feedback transforms simple stochastic switching into a rich dynamical system with memory, non‑trivial timing statistics, and inter‑switch correlations. The results are directly testable experimentally (e.g., via time‑lapse fluorescence microscopy of reporter constructs) and lay a solid foundation for exploring more complex feedback topologies, spatial effects, and multi‑enzyme networks in future work.
Comments & Academic Discussion
Loading comments...
Leave a Comment