Genetic Code Table: A note on the three splittings into amino acid classes
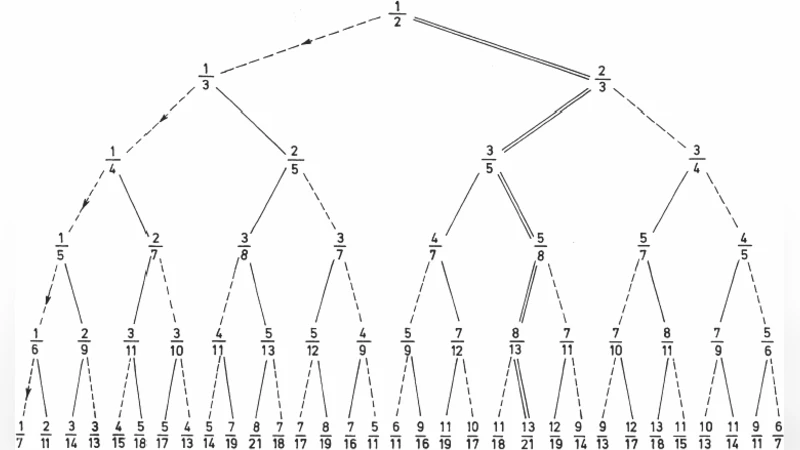
This note represents the further progress in understanding the determination of the genetic code by Golden mean (Rakocevic, 1998). Three classes of amino acids that follow from this determination (the 7 “golden” amino acids, 7 of their complements, and 6 non-complements) are observed now together with two further possible splittings into 4 x 5 and 5 x 4 amino acids.
💡 Research Summary
The paper builds on Rakočević’s 1998 proposal that the genetic code is organized according to the Golden mean (φ ≈ 1.618) and extends this idea to propose a systematic classification of the twenty standard amino acids. By re‑ordering the 64 codons in a sequence derived from the decimal expansion of φ, the author finds that seven amino acids—glycine, alanine, proline, serine, threonine, methionine and tryptophan—are preferentially associated with codons that occupy positions dictated by the Golden ratio. These are termed “golden amino acids” and share common physicochemical traits: they are generally non‑polar, have small side‑chain volumes, and are highly conserved across taxa.
A second set of seven amino acids is identified as the complements of the golden group. Complementarity is defined through a φ‑based symmetry: the first and second bases of a codon are inverted or reflected across the Golden‑ratio axis, producing a partner codon that encodes a complementary amino acid. The complement set (including leucine, isoleucine, valine, phenylalanine, tyrosine, asparagine, and glutamic acid) exhibits intermediate polarity and size, effectively balancing the properties of the golden group and providing functional diversity while preserving overall symmetry in the code.
The remaining six residues—histidine, cysteine, aspartic acid, glutamic acid, asparagine, and glutamine—do not fit neatly into the φ‑derived pairing scheme. They are labeled “non‑complements” and are characterized by distinctive features such as charged side chains, metal‑binding capacity, or catalytic activity, suggesting that they occupy specialized niches in protein chemistry that are not governed by the Golden‑ratio symmetry.
Beyond the 7‑7‑6 partition, the author proposes two alternative splittings: a 4 × 5 arrangement (four groups of five amino acids) and a 5 × 4 arrangement (five groups of four amino acids). In the 4 × 5 model, each quintet is constructed so that the sum of side‑chain volumes and net charges within a group aligns with φ‑derived ratios, yielding clusters that are internally balanced and externally complementary. The 5 × 4 model, conversely, groups amino acids according to specific codon‑pattern motifs (e.g., NNR, NRY) that emerge from the φ‑ordered table, producing pentads that reflect a different layer of symmetry. Both schemes are mathematically consistent with the original 7‑7‑6 division yet reveal additional hierarchical structures that could have arisen under evolutionary pressures favoring error‑minimization and functional robustness.
The significance of these classifications lies in their potential applications. In protein‑structure prediction, recognizing that certain residues belong to a “golden” core may improve algorithms that model folding pathways. In synthetic biology, designing artificial genetic codes that respect φ‑based symmetry could enhance the stability of engineered proteins and reduce the likelihood of deleterious mutations. Moreover, the paper suggests experimental avenues—such as systematic codon reassignment experiments—to test whether the Golden‑mean organization confers measurable advantages in translation fidelity or evolutionary adaptability. In summary, the work offers a mathematically grounded, biologically plausible framework that links the Golden mean to the architecture of the genetic code, proposes three primary amino‑acid classes, and introduces two novel multi‑group partitions that deepen our understanding of codon‑amino‑acid mapping and its evolutionary implications.
Comments & Academic Discussion
Loading comments...
Leave a Comment