Cortical Factor Feedback Model for Cellular Locomotion and Cytofission
Eukaryotic cells can move spontaneously without being guided by external cues. For such spontaneous movements, a variety of different modes have been observed, including the amoeboid-like locomotion with protrusion of multiple pseudopods, the keratocyte-like locomotion with a widely spread lamellipodium, cell division with two daughter cells crawling in opposite directions, and fragmentations of a cell to multiple pieces. Mutagenesis studies have revealed that cells exhibit these modes depending on which genes are deficient, suggesting that seemingly different modes are the manifestation of a common mechanism to regulate cell motion. In this paper, we propose a hypothesis that the positive feedback mechanism working through the inhomogeneous distribution of regulatory proteins underlies this variety of cell locomotion and cytofission. In this hypothesis, a set of regulatory proteins, which we call cortical factors, suppress actin polymerization. These suppressing factors are diluted at the extending front and accumulated at the retracting rear of cell, which establishes a cellular polarity and enhances the cell motility, leading to the further accumulation of cortical factors at the rear. Stochastic simulation of cell movement shows that the positive feedback mechanism of cortical factors stabilizes or destabilizes modes of movement and determines the cell migration pattern. The model predicts that the pattern is selected by changing the rate of formation of the actin-filament network or the threshold to initiate the network formation.
💡 Research Summary
The paper proposes a unifying hypothesis that a single positive‑feedback loop, mediated by “cortical factors” – a set of regulatory proteins that suppress actin polymerization – can generate the wide spectrum of spontaneous cell motility modes and cytofission observed in eukaryotes. In the model, cortical factors become locally diluted at the protruding front of a cell while they accumulate at the retracting rear. This spatial inhomogeneity creates polarity: reduced inhibition at the front permits rapid actin filament nucleation and membrane extension, whereas increased inhibition at the rear reinforces contractility and retraction. The resulting feedback amplifies the initial polarity, stabilizing the direction of movement and further sharpening the rear‑front gradient of cortical factors.
To test the idea, the authors built a stochastic, lattice‑based simulation in which each lattice site tracks the concentration of cortical factors, the density of actin filaments, and the local membrane state. The dynamics are governed primarily by two tunable parameters: (i) the rate of actin‑network formation (k_form) and (ii) the threshold concentration of cortical factors required to trigger network formation (θ_act). By varying these parameters the model reproduces four characteristic behaviors that have been documented experimentally:
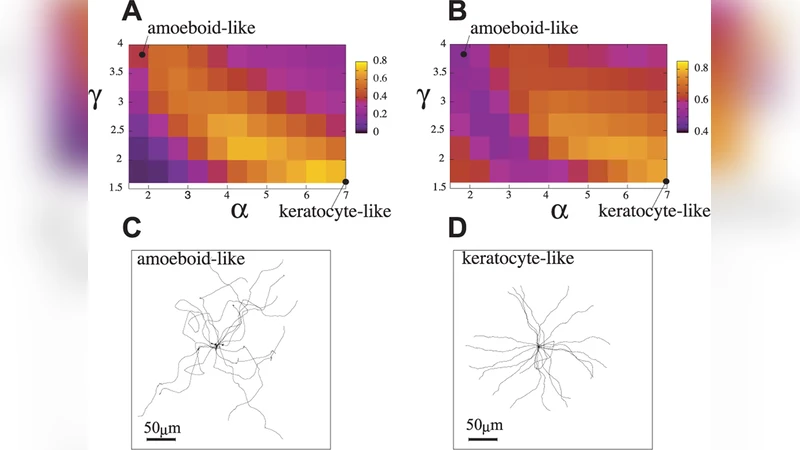
- Keratinocyte‑like migration – high k_form combined with low θ_act yields a broad, stable lamellipodium that spreads laterally, producing smooth, persistent forward motion.
- Amoeboid‑like migration – low k_form and high θ_act generate intermittent, localized actin bursts that give rise to multiple, transient pseudopods and a highly exploratory trajectory.
- Cytofission – intermediate values cause the cell to split its leading edge into two symmetric fronts that move in opposite directions, mimicking the division‑like crawling of daughter cells observed in certain mutants.
- Fragmentation (cytofission‑like fission) – excessive accumulation of cortical factors at the rear creates a strong contractile “pinch” that fragments the cell into several pieces.
Crucially, all these modes emerge from the same underlying feedback circuit; only the kinetic parameters governing actin assembly and cortical‑factor thresholds are altered. This provides a mechanistic explanation for why diverse genetic knock‑outs (e.g., loss of myosin II, Arp2/3 complex components, or regulators of phosphoinositide signaling) can each produce a distinct motility phenotype while still being rooted in a common regulatory architecture.
The simulation makes several experimentally testable predictions. Pharmacological inhibition of myosin II, which would reduce rear‑ward cortical‑factor accumulation, should shift cells from a fragmented or cytofission phenotype toward a broader lamellipodial spread. Conversely, drugs that hyper‑activate the Arp2/3 complex (thereby increasing k_form) are predicted to suppress multiple pseudopod formation and promote smoother, keratocyte‑type migration. The model also predicts that modest changes in the cortical‑factor diffusion coefficient can tip the system from a stable to an unstable regime, leading to spontaneous polarity reversals.
In the discussion, the authors argue that the cortical‑factor feedback loop offers a parsimonious framework for integrating cell‑intrinsic cues (such as internal protein concentrations) with external perturbations (chemical inhibitors, substrate stiffness) to control cell shape, speed, and directionality. They suggest that targeting components of this loop could provide new strategies for controlling pathological cell migration, for example, limiting metastatic cancer cell invasion or modulating immune‑cell trafficking during inflammation.
Overall, the study combines theoretical modeling with biologically realistic parameters to show that a simple positive‑feedback mechanism based on the spatial redistribution of actin‑suppressing factors can account for the diversity of spontaneous locomotion and cytofission phenotypes. It bridges the gap between genetic observations and physical‑chemical principles of cell motility, offering a fertile ground for future experimental validation and therapeutic exploration.
Comments & Academic Discussion
Loading comments...
Leave a Comment