Lethal Mutagenesis in Viruses and Bacteria
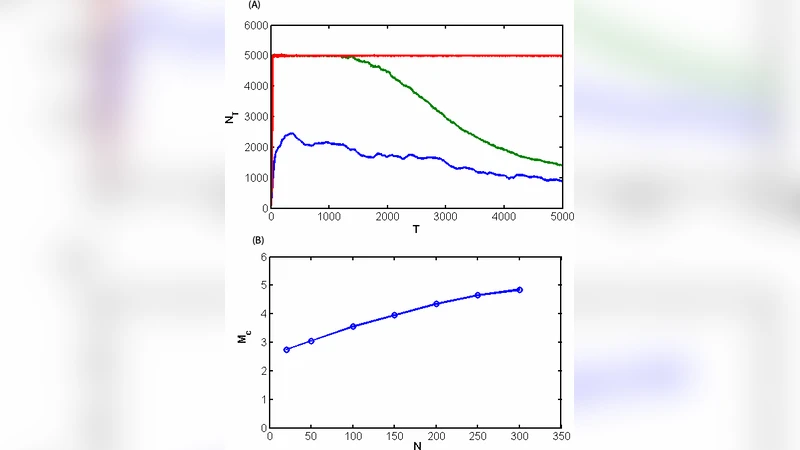
Here we study how mutations which change physical properties of cell proteins (stability) impact population survival and growth. In our model the genotype is presented as a set of N numbers, folding free energies of cells N proteins. Mutations occur upon replications so that stabilities of some proteins in daughter cells differ from those in parent cell by random amounts drawn from experimental distribution of mutational effects on protein stability. The genotype-phenotype relationship posits that unstable proteins confer lethal phenotype to a cell and in addition the cells fitness (duplication rate) is proportional to the concentration of its folded proteins. Simulations reveal that lethal mutagenesis occurs at mutation rates close to 7 mutations per genome per replications for RNA viruses and about half of that for DNA based organisms, in accord with earlier predictions from analytical theory and experiment. This number appears somewhat dependent on the number of genes in the organisms and natural death rate. Further, our model reproduces the distribution of stabilities of natural proteins in excellent agreement with experiment. Our model predicts that species with high mutation rates, tend to have less stable proteins compared to species with low mutation rate.
💡 Research Summary
The paper presents a quantitative, physics‑based model that links genetic mutations to the thermodynamic stability of cellular proteins and, through this link, to the survival and replication dynamics of viral and bacterial populations. Each organism’s genotype is represented as a vector of N folding free energies (ΔG) corresponding to its N essential proteins. During replication, mutations occur with a per‑genome rate u; each mutation changes the ΔG of the affected protein by a random amount drawn from an experimentally measured distribution of mutational effects on protein stability (ΔΔG), typically a near‑Gaussian with mean ≈0 and σ ≈ 1 kcal mol⁻¹.
The genotype‑phenotype map is deliberately simple yet biologically grounded. If any protein in a cell becomes thermodynamically unstable (ΔG > 0), the cell is assigned a lethal phenotype and is removed from the population. If all proteins remain stable (ΔG < 0), the cell’s replication rate r is proportional to the product of the folded fractions of its proteins. Mathematically, r = r₀ ∏₁ᴺ
Comments & Academic Discussion
Loading comments...
Leave a Comment