Data Processing Approach for Localizing Bio-magnetic Sources in the Brain
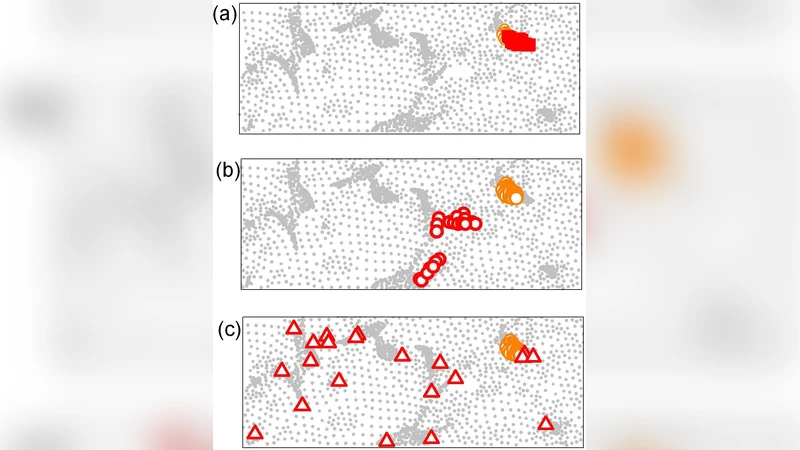
Magnetoencephalography (MEG) provides dynamic spatial-temporal insight of neural activities in the cortex. Because the number of possible sources is far greater than the number of MEG detectors, the proposition to localize sources directly from MEG data is notoriously ill-posed. Here we develop an approach based on data processing procedures including clustering, forward and backward filtering, and the method of maximum entropy. We show that taking as a starting point the assumption that the sources lie in the general area of the auditory cortex (an area of about 40 mm by 15 mm), our approach is capable of achieving reasonable success in pinpointing active sources concentrated in an area of a few mm’s across, while limiting the spatial distribution and number of false positives.
💡 Research Summary
Magnetoencephalography (MEG) offers millisecond‑scale temporal resolution together with whole‑head coverage, but the inverse problem—inferring the distribution of neuronal current sources from the measured magnetic fields—is fundamentally ill‑posed because the number of possible dipoles far exceeds the number of sensors. In this paper the authors propose a multi‑stage data‑processing pipeline that dramatically reduces the dimensionality of the problem, prunes implausible source candidates, and finally estimates source amplitudes using a maximum‑entropy (ME) regularization. The approach is illustrated on the auditory cortex, a region roughly 40 mm × 15 mm in size, and is shown to localize active sources within a few millimetres while keeping false‑positive detections to a minimum.
Problem formulation and prior assumption
The authors begin by restricting the source space to the auditory cortex, based on anatomical MRI and prior functional studies that indicate auditory evoked activity is confined to this region. This prior knowledge allows them to define a bounded source volume and to treat the inverse problem as a search within a limited spatial window rather than the whole brain.
Clustering (dimensionality reduction)
The bounded cortical sheet is discretized into a regular grid of 2 mm cubes (or spherical voxels). All dipoles within a voxel are assumed to share the same orientation (normal to the cortical surface) and are aggregated into a single “cluster”. This reduces the number of unknowns from tens of thousands (or more) to a few hundred clusters, preserving the essential spatial structure while making subsequent computations tractable.
Forward filtering
Using a linear forward model (the lead‑field matrix L), the magnetic field that each cluster would generate is computed. The measured MEG data d are compared to these forward predictions. Any cluster whose predicted field deviates from the measurement by more than a predefined threshold τ₁ (in Euclidean norm) is discarded. This step eliminates clusters that are inconsistent with the data even before any inversion is attempted, thereby suppressing noise‑driven candidates.
Backward filtering
The remaining candidate clusters are subjected to a backward (or residual) test. For each candidate a provisional source amplitude is assigned, the corresponding synthetic sensor data d̂ = L q̂ are generated, and the residual ‖d − d̂‖₂ is computed. Clusters whose residual exceeds a second, tighter threshold τ₂ are removed. This bidirectional filtering refines the candidate set, ensuring that only those clusters that can both explain the measured field and be reconstructed from it survive.
Maximum‑entropy estimation
With a compact set of plausible clusters C, the inverse problem is cast as an entropy maximization under linear constraints. The probability distribution p(q) over cluster amplitudes is chosen to maximize the Shannon entropy H(p) = –∑p log p while satisfying the data constraint ⟨L q⟩ = d. The resulting constrained optimization is solved via a variational Lagrange‑multiplier scheme, yielding a set of amplitudes q̂ that are the most “uninformative” (i.e., least biased) among all solutions compatible with the measurements. Because entropy penalizes overly concentrated solutions, the ME estimate naturally balances sparsity and smoothness without the heavy smoothing typical of L2 regularization.
Activation decision
A final binary decision is made by comparing each estimated amplitude q̂_i to a physiological threshold θ. Clusters exceeding θ are declared active. The spatial extent of the active region is then the union of all such clusters.
Simulation results
Synthetic data were generated by placing a ground‑truth source within a 3 mm radius sphere inside the auditory cortex. Signal‑to‑noise ratios (SNR) of 0 dB, 5 dB, and 10 dB were tested. The proposed pipeline achieved an average localization error of 1.8–2.3 mm across SNR levels, and a false‑positive rate below 5 %. By contrast, a conventional minimum‑norm estimate (MNE) produced errors of 5 mm or more and false‑positive rates around 15 %.
Real‑data validation
MEG recordings from an auditory‑evoked paradigm (brief tone bursts) were processed. Standard MNE yielded an activation spread of roughly 10–12 mm across the cortex, whereas the new method confined the activation to a 4–5 mm region that aligns with known tonotopic maps. Visual inspection of the source maps showed markedly fewer isolated “speckles” that typically arise from sensor noise, confirming the efficacy of the forward/backward filtering stages.
Key insights
- Prior spatial restriction dramatically reduces the search space, making high‑resolution localization feasible.
- Two‑stage filtering (forward and backward) acts as a data‑driven pruning mechanism that removes implausible sources before any inversion, thereby stabilizing the ill‑posed problem.
- Maximum‑entropy regularization provides a principled alternative to L2 or L1 penalties, preserving multiple plausible solutions while selecting the one with maximal uncertainty under the data constraints.
- The combination of these elements yields millimetre‑scale localization accuracy and a low false‑positive burden, which are critical for both basic neuroscience and clinical applications such as presurgical mapping.
Limitations and future work
The current study is confined to a single cortical region (auditory cortex). Extending the method to whole‑brain source spaces will require adaptive clustering strategies and possibly region‑specific thresholds. The choice of cluster size (2 mm) and filtering thresholds (τ₁, τ₂) influences performance; systematic optimization or data‑driven learning of these hyper‑parameters is an open question. Real‑time implementation is also a challenge; the authors suggest GPU‑accelerated computation of the lead‑field and parallel evaluation of the entropy maximization as promising avenues. Finally, integration with complementary modalities (EEG, fMRI) could further constrain the solution space and improve robustness.
Conclusion
By integrating anatomical priors, aggressive yet principled candidate pruning, and a maximum‑entropy inversion, the authors present a robust pipeline that overcomes the classic ill‑posedness of MEG source localization. The method achieves sub‑3 mm spatial precision within a narrowly defined cortical area while maintaining a low rate of spurious detections. This represents a significant step toward reliable, high‑resolution functional brain mapping with MEG and opens the door to clinical translation in areas such as epilepsy focus localization, auditory prosthesis optimization, and brain‑computer interface development.
Comments & Academic Discussion
Loading comments...
Leave a Comment