In this paper, a model for understanding the effects of selection using systems- level computational approaches is introduced. A number of concepts and principles essential for understanding the motivation for constructing the model will be introduced first. This will be followed by a description of parameters, measurements, and graphical representations used in the model. Four possible outcomes for this model are then introduced and described. In addition, the relationship of relative fitness to selection is described. Finally, the consequences and potential lessons learned from the model are discussed.
Deep Dive into Hierarchies of Biocomplexity: modeling lifes energetic complexity.
In this paper, a model for understanding the effects of selection using systems- level computational approaches is introduced. A number of concepts and principles essential for understanding the motivation for constructing the model will be introduced first. This will be followed by a description of parameters, measurements, and graphical representations used in the model. Four possible outcomes for this model are then introduced and described. In addition, the relationship of relative fitness to selection is described. Finally, the consequences and potential lessons learned from the model are discussed.
Hierarchies of Biocomplexity: modeling life’s energetic complexity
Bradly Alicea1
1MIND Lab, Michigan State University, East Lansing, MI. USA, 48824
E-mail: freejumper@yahoo.com
Abstract. In this paper, a model for understanding the effects of selection using systems-level
computational approaches is introduced. A number of concepts and principles essential for
understanding the motivation for constructing the model will be introduced first. This will be followed
by a description of parameters, measurements, and graphical representations used in the model. Four
possible outcomes for this model are then introduced and described. In addition, the relationship of
relative fitness to selection is described. Finally, the consequences and potential lessons learned from
the model are discussed.
Introduction.
In this paper, a computational approach to biological complexity at the level of a single organism is
introduced. This model might be applied to a number of domains. For example, such a model might be
applied to the field of synthetic biology [1]. Currently, most synthetic biology has focused on genomic
function, whereas this model would allow for a better understanding of physiological regulation.
Another potential application domain is in the area of programmable artificial muscle [2], particularly
in the implementation of actuators with adaptive contraction. Thirdly, this model may serve as a general
model for applying computational methods to medical problems such as the complex physiology of
disease states.
The theory of evolution depends on natural selection and neutral processes, which in turn depend
upon interactions at multiple scales of analysis. As a secondary aim, I will take the position that the
effects of environmental selection on organismal1 physiology can be abstracted to a series of units that
replicate at a specific level and are interconnected with units at other levels. This relates to outstanding
controversies in multilevel selection, as a classic problem is approached using systems-level and
computational approaches.
1.1 Introduction to problem.
Variation is the raw material of evolution. In biological systems, this variation is ubiquitous, existing
at every scale of analysis. It can also be argued that the self-organization of biocomplexity is the
product of two phenomena: natural selection and the existence of multi-scalar phenomena. To better
understand how these phenomena are central to determining the dynamics of self-organization, the
levels of selection problem and the concept of hierarchical energetic organization will be brought to
bear on the problem.
1.2 Levels of selection.
While the levels of selection problem is a controversial topic, the idea of selection acting at multiple
levels of biological organization can be informative for understanding the structure of biocomplexity.
Selection can act at multiple levels, from molecular to phenotypic to cultural [3]. When selection acting
at one scale is observed at another scale, its effects may appear to be diffuse. The magnitude of this
effect is dependent on the units of selection at a certain level. Dawkins [4] conceives of these units as
replicators. These replicator units will become important in understanding the structure of each level
and interactions between levels.
1 Organismal can be defined as having to do with the properties of the organism.
1.3 Introduction to scale.
Intuitively, a difference in scale refers to an order of magnitude. The concept of scale in
biocomplexity has been explored using a number of different examples and perspectives [5, 6]. Most
recently, this approach has been applied to understanding physiological phenomena [7]. In the
abstraction presented here, a single level can be better understood as a set of scalar elements (see
Figure 1) with two properties: 1) the elements of a single scalar set are embedded in a hierarchy of
scalar sets, and 2) each scalar set contains multiple replicators.
Thinking of the levels of selection problem in this way may resolve a central contention in modern
conceptualizations of multi-level selection: how does selection at a particular scale affect features at
other scales? In other words, how does group selection affect the relative fitness of an organism, or how
do changes in gene expression and phenotype interact? These questions are especially important when
we consider that elements at a lower scale (e.g cell populations) interact with each other to form
emergent properties at higher scales (e.g. organs).
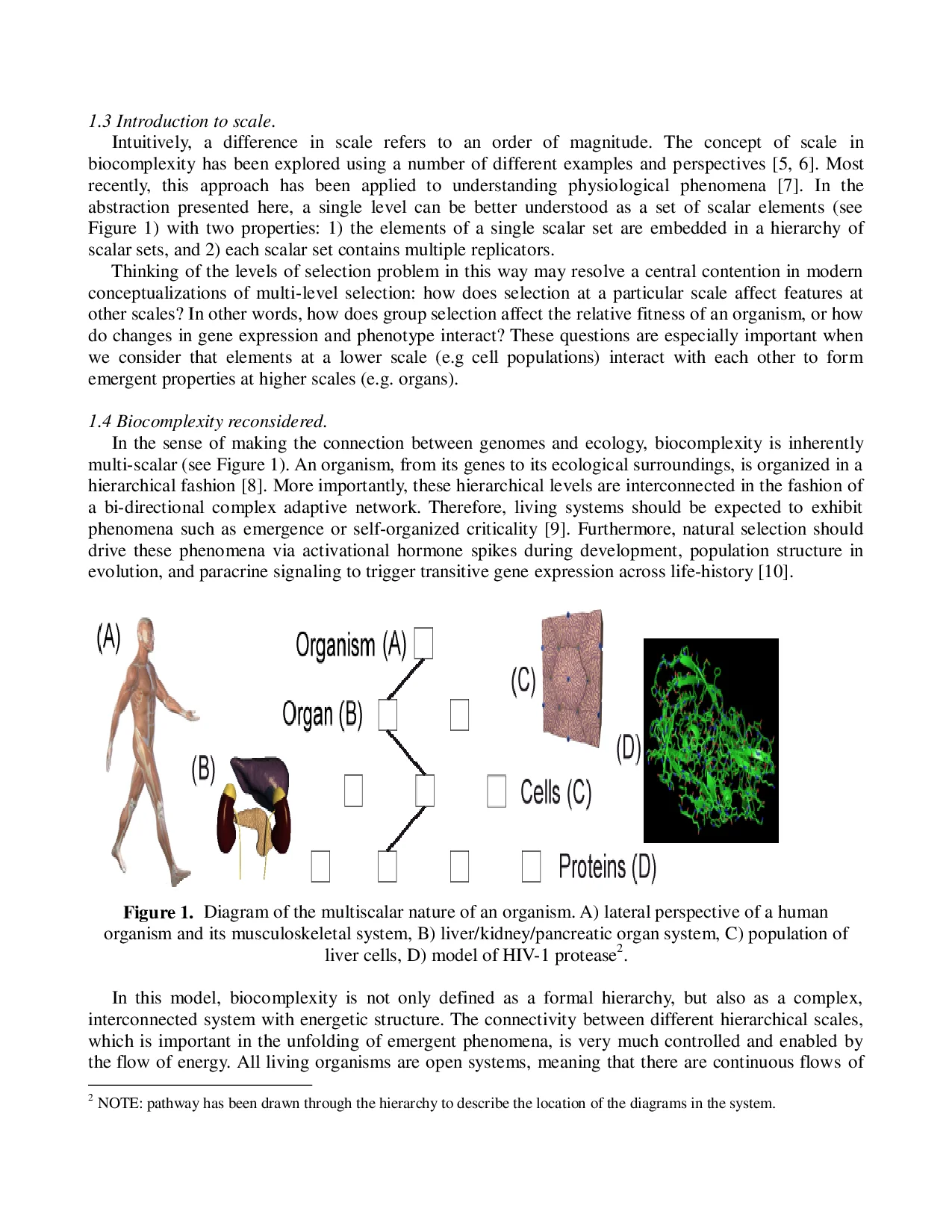
1.4 Biocomplexity reconsidered.
In the sense of making the connection between genomes and ecology, biocomplexity is inherently
multi-scalar (see Figure 1). An organism, from its genes to its ecological surroundings, is organized in a
hierarchical fashion [8]. More importantly, these hierarchical le
…(Full text truncated)…
This content is AI-processed based on ArXiv data.