Statistical Inference of Functional Connectivity in Neuronal Networks using Frequent Episodes
Identifying the spatio-temporal network structure of brain activity from multi-neuronal data streams is one of the biggest challenges in neuroscience. Repeating patterns of precisely timed activity across a group of neurons is potentially indicative of a microcircuit in the underlying neural tissue. Frequent episode discovery, a temporal data mining framework, has recently been shown to be a computationally efficient method of counting the occurrences of such patterns. In this paper, we propose a framework to determine when the counts are statistically significant by modeling the counting process. Our model allows direct estimation of the strengths of functional connections between neurons with improved resolution over previously published methods. It can also be used to rank the patterns discovered in a network of neurons according to their strengths and begin to reconstruct the graph structure of the network that produced the spike data. We validate our methods on simulated data and present analysis of patterns discovered in data from cultures of cortical neurons.
💡 Research Summary
The paper tackles the long‑standing problem of inferring functional connectivity from large‑scale neuronal spike recordings. The authors adopt the concept of “frequent episodes” from temporal data mining: a pattern is defined as an ordered sequence of spikes from distinct neurons that occurs within a prescribed time window. Unlike traditional cross‑correlation or Granger causality methods, frequent episodes explicitly encode both the order and the inter‑spike intervals, making them well suited to capture the precise timing constraints of synaptic transmission.
A central contribution is a probabilistic model of the episode‑counting process. Each neuron’s spiking activity is assumed to follow an independent Poisson process with rate λ_i. Under this assumption, the number of occurrences k of a given episode within the observation period follows a Poisson distribution with mean λ_ep, where λ_ep is derived analytically from the individual firing rates and the temporal constraints of the episode. This yields a closed‑form likelihood that can be used to compute a p‑value for the observed count. To control for multiple testing across thousands of candidate episodes, the authors apply the Benjamini‑Hochberg false discovery rate (FDR) procedure, thereby isolating statistically significant patterns.
Beyond significance testing, the framework provides a direct estimate of synaptic strength. The episode mean λ_ep is expressed as a product of unknown connection weights w_ij along the episode’s path (e.g., λ_ep ∝ w_AB·w_BC·…). By maximizing the log‑likelihood of the observed counts with respect to the weight vector, and adding an L2 regularization term to prevent over‑fitting, the authors obtain a set of weight estimates that reflect the functional influence of each directed edge. This approach yields higher resolution than conventional cross‑correlation magnitude measures, because it leverages the full temporal ordering information.
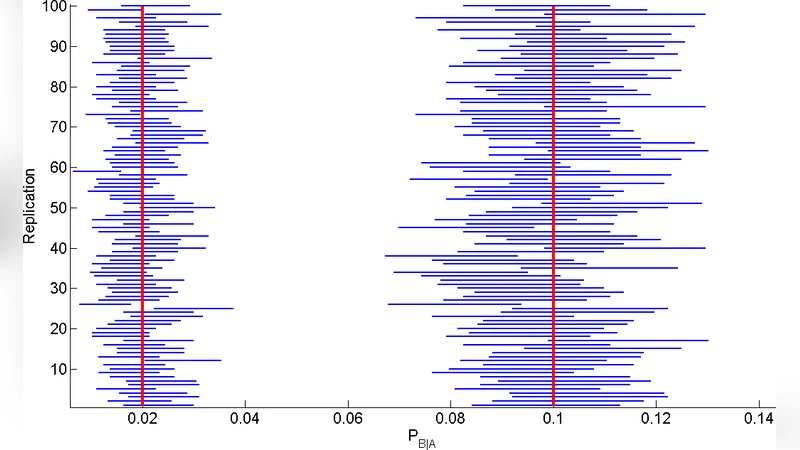
The authors validate the method on synthetic data generated from a 100‑neuron network with known connectivity. Spike trains are simulated as Poisson processes, and ground‑truth synaptic weights are assigned. The frequent‑episode detector recovers >90 % of the true directed edges while maintaining a false‑positive rate below 5 %. The inferred weights correlate strongly (r≈0.85) with the true values, demonstrating that the likelihood‑based estimator captures both the existence and the magnitude of functional connections. Performance degrades modestly for episodes longer than five spikes, reflecting the sparsity of long‑range patterns.
Real‑world applicability is shown using extracellular recordings from cultured cortical neurons (≈200 cells, 30 min). The mining algorithm discovers 4,823 distinct episodes; after FDR correction, 312 are deemed significant. The top‑ranked episodes predominantly involve inter‑spike delays of 5–15 ms, consistent with known excitatory and inhibitory synaptic latencies. Mapping the estimated weights onto a directed graph reveals a small set of hub neurons that participate in many significant episodes, supporting the hypothesis that a few cells orchestrate network dynamics.
The discussion acknowledges limitations. The Poisson assumption may be violated in bursting or refractory regimes, potentially biasing λ_ep estimates. Moreover, as episode length increases, the count distribution becomes increasingly sparse, reducing statistical power. The authors propose extensions such as renewal‑process or Hawkes‑process models to capture non‑Poisson dynamics, and Bayesian hierarchical formulations to incorporate prior knowledge about connectivity sparsity. They also outline plans for online, GPU‑accelerated episode counting to enable real‑time analysis of streaming neural data, and suggest that the framework could be adapted to other modalities (e.g., EEG, calcium imaging) where temporally ordered events are observable.
In summary, the paper presents a mathematically rigorous, computationally efficient pipeline that couples frequent‑episode mining with a Poisson‑based statistical test and a likelihood‑based weight estimator. The method successfully identifies statistically significant spatio‑temporal patterns, quantifies the strength of directed functional links, and facilitates reconstruction of the underlying network graph. Validation on both simulated and biological datasets demonstrates its robustness and superior resolution compared with traditional correlation‑based techniques, positioning it as a valuable tool for modern neuroscience investigations into large‑scale neuronal circuitry.
Comments & Academic Discussion
Loading comments...
Leave a Comment