Conformational Transitions in Molecular Systems
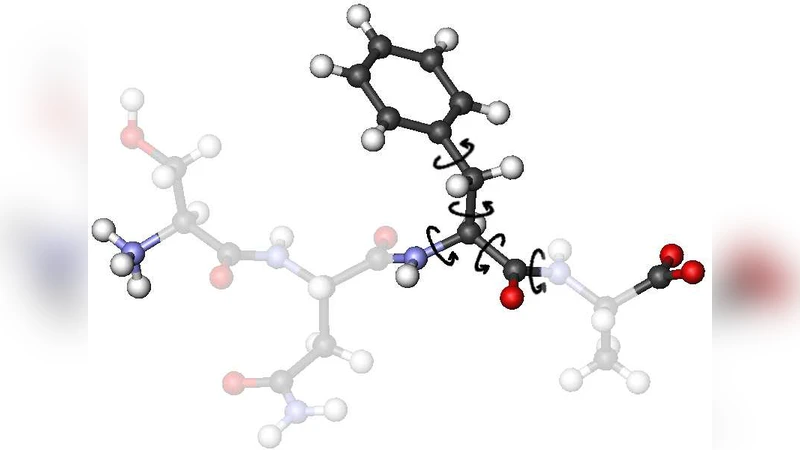
Proteins are the “work horses” in biological systems. In almost all functions specific proteins are involved. They control molecular transport processes, stabilize the cell structure, enzymatically catalyze chemical reactions; others act as molecular motors in the complex machinery of molecular synthetization processes. Due to their significance, misfolds and malfunctions of proteins typically entail disastrous diseases, such as Alzheimer’s disease and bovine spongiform encephalopathy (BSE). Therefore, the understanding of the trinity of amino acid composition, geometric structure, and biological function is one of the most essential challenges for the natural sciences. Here, we glance at conformational transitions accompanying the structure formation in protein folding processes.
💡 Research Summary
**
The paper addresses the fundamental problem of how proteins acquire their functional three‑dimensional structures, focusing on the conformational transitions that occur during folding. After outlining the diverse biological roles of proteins—transport, structural support, catalysis, and molecular motors—the authors frame protein folding as a navigation of a rugged energy landscape. In this landscape, numerous local minima correspond to partially folded intermediates, while the high‑energy barriers separating them define transition states. These transition states are fleeting, structurally heterogeneous configurations that dictate the folding pathway and its kinetic bottlenecks.
Experimental techniques such as Φ‑value analysis, NMR relaxation dispersion, and single‑molecule force spectroscopy are reviewed for their ability to capture transition‑state ensembles. Complementarily, the authors discuss advanced computational approaches, notably molecular dynamics simulations enhanced by adaptive sampling and reinforcement‑learning algorithms, which allow efficient exploration of high‑dimensional conformational space and identification of minimum‑energy paths.
A central theme is the role of “core contacts” and “core residues” in shaping the transition‑state barrier. Core contacts—hydrophobic packing, hydrogen bonds, and electrostatic interactions—form the scaffold of the transition state, while mutations in core residues can dramatically raise or lower the barrier. The paper illustrates this with case studies of amyloid‑β and prion proteins, showing how disease‑associated mutations reroute folding pathways toward off‑pathway aggregates that underlie Alzheimer’s disease and BSE.
The authors then explore therapeutic implications. Transition‑state inhibitors are proposed as a class of molecules that either stabilize the native transition state, thereby smoothing the folding funnel, or increase the barrier of pathogenic off‑pathway states, preventing toxic aggregation. A workflow combining structure‑based virtual screening, free‑energy perturbation calculations, and machine‑learning predictors is presented as a blueprint for rational drug design targeting folding transitions.
In conclusion, the paper argues that an integrated experimental‑computational strategy is essential for a quantitative understanding of protein folding transitions. Such insight not only clarifies the mechanistic link between amino‑acid sequence, structure, and function but also opens avenues for intervening in misfolding diseases through transition‑state‑focused therapeutics. Future directions include ultrafast single‑molecule measurements, QM/MM hybrid simulations, and large‑scale AI models to predict transition pathways across the proteome.
Comments & Academic Discussion
Loading comments...
Leave a Comment