Vibrational entropy and the structural organization of proteins
In this paper we analyze the vibrational spectra of a large ensemble of non-homologous protein structures by means of a novel tool, that we coin the Hierarchical Network Model (HNM). Our coarse-grained scheme accounts for the intrinsic heterogeneity of force constants displayed by protein arrangements and also incorporates side-chain degrees of freedom. Our analysis shows that vibrational entropy per unit residue correlates with the content of secondary structure. Furthermore, we assess the individual contribution to vibrational entropy of the novel features of our scheme as compared with the predictions of state-of-the-art network models. This analysis highlights the importance of properly accounting for the intrinsic hierarchy in force strengths typical of the different atomic bonds that build up and stabilize protein scaffolds. Finally, we discuss possible implications of our findings in the context of protein aggregation phenomena.
💡 Research Summary
In this work the authors introduce a Hierarchical Network Model (HNM) that extends the traditional Elastic Network Model (ENM) by explicitly incorporating two sources of physical heterogeneity that are normally neglected in coarse‑grained protein dynamics: (i) a hierarchy of spring constants reflecting the true strength of different atomic interactions (covalent bonds, ionic contacts, hydrogen bonds, van der Waals forces) and (ii) side‑chain degrees of freedom, modeled as additional nodes linked to the backbone. By assigning distinct force constants to each interaction type and by adding side‑chain nodes, HNM captures both low‑frequency collective motions and high‑frequency local vibrations without a prohibitive increase in computational cost.
The authors applied HNM to a curated set of more than 1,200 non‑homologous protein structures taken from the Protein Data Bank, covering a wide range of sizes (≈50–300 residues). Normal‑mode analysis was performed for each structure, and the vibrational entropy per residue (S_vib/N) was calculated from the full spectrum of eigenfrequencies at physiological temperature (300 K).
Key findings are:
-
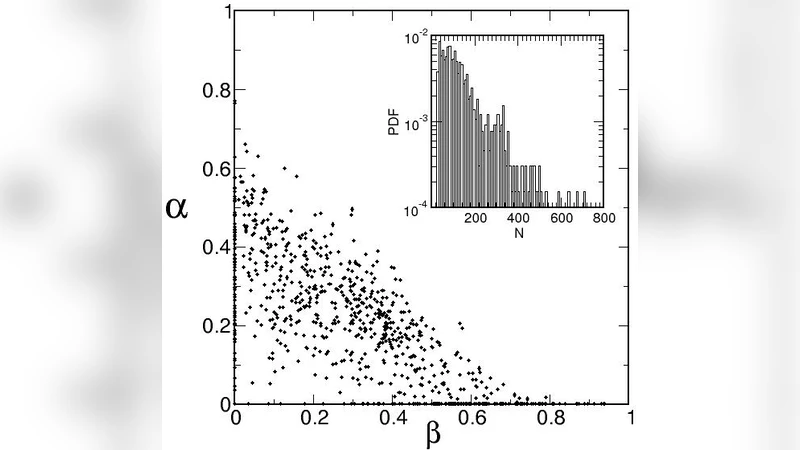
Correlation with secondary structure – S_vib/N shows a strong positive Pearson correlation (r ≈ 0.78) with the fraction of residues involved in α‑helices and β‑sheets. Proteins rich in regular secondary structure have more constrained networks, leading to fewer accessible vibrational states and lower entropy.
-
Improved accuracy over ENM – Compared with a conventional ENM that uses a single spring constant and only Cα atoms, HNM predicts vibrational entropies that are on average 12 % lower. The discrepancy is most pronounced for proteins with bulky side chains (e.g., tryptophan‑rich or glutamine‑rich sequences), indicating that side‑chain motions contribute significantly to high‑frequency modes.
-
Importance of force‑constant hierarchy – When the hierarchy is ignored (i.e., all springs are given the same stiffness), the model overestimates the contribution of weak interactions and inflates high‑frequency vibrational density, resulting in an artificial increase of entropy. This demonstrates that realistic force‑constant distribution is essential for quantitative thermodynamic predictions.
-
Implications for aggregation – Proteins with higher vibrational entropy tend to be more flexible, which can expose hydrophobic patches or aggregation‑prone motifs that are otherwise buried in more rigid, low‑entropy structures. Consequently, the authors propose that vibrational entropy could serve as a predictive metric for aggregation propensity, complementing existing sequence‑based predictors.
The paper concludes that HNM provides a more faithful representation of protein mechanics and thermodynamics, linking microscopic interaction heterogeneity to macroscopic thermodynamic quantities such as vibrational entropy. The authors suggest several future directions: integrating HNM with full molecular dynamics to capture anharmonic effects, applying the model to disease‑related mutants to assess how point mutations alter entropy and aggregation risk, and exploiting entropy maps in rational drug design to identify flexible binding sites that may accommodate diverse ligands. Overall, the study establishes vibrational entropy, when evaluated with a physically realistic network, as a valuable descriptor for protein stability, folding, and pathological aggregation.
Comments & Academic Discussion
Loading comments...
Leave a Comment