Embedding Data within Knowledge Spaces
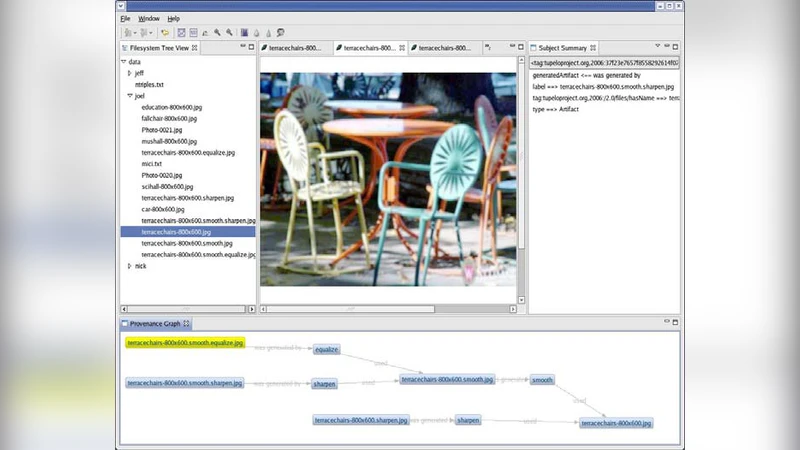
The promise of e-Science will only be realized when data is discoverable, accessible, and comprehensible within distributed teams, across disciplines, and over the long-term–without reliance on out-of-band (non-digital) means. We have developed the open-source Tupelo semantic content management framework and are employing it to manage a wide range of e-Science entities (including data, documents, workflows, people, and projects) and a broad range of metadata (including provenance, social networks, geospatial relationships, temporal relations, and domain descriptions). Tupelo couples the use of global identifiers and resource description framework (RDF) statements with an aggregatable content repository model to provide a unified space for securely managing distributed heterogeneous content and relationships.
💡 Research Summary
The paper presents Tupelo, an open‑source semantic content management framework designed to meet the fundamental requirements of e‑Science: data must be discoverable, accessible, and understandable across distributed teams, multiple disciplines, and long time spans without reliance on non‑digital mechanisms. The authors argue that traditional file‑system or database‑centric solutions fall short because they separate the physical storage of data from its descriptive metadata, leading to fragmented provenance, inconsistent access control, and difficulties in long‑term preservation.
Tupelo’s architecture is built around three core principles. First, every entity—whether a raw data file, a document, a workflow, a person, or a project—is assigned a globally unique identifier (URI). Relationships among these entities are expressed as RDF triples, enabling a graph‑based representation of complex dependencies such as provenance chains, social networks, geospatial links, and temporal ordering. Second, the framework adopts an “aggregatable content repository” model. It abstracts away the heterogeneity of underlying storage back‑ends (local file systems, cloud object stores, relational databases, etc.) by providing a uniform API that treats all resources as part of a single logical repository. Third, security and policy enforcement are integrated at the semantic level: access control rules can be attached to individual triples, allowing fine‑grained permissions that cover both the binary payload and its associated metadata.
Implementation-wise, Tupelo couples an RDF store (Apache Jena TDB or Blazegraph) with a Java Content Repository (JCR) implementation (Apache Jackrabbit). The RDF layer handles all metadata, supporting SPARQL queries and a RESTful SPARQL endpoint for programmatic access. The JCR layer stores the actual binary content, offering versioning, check‑in/check‑out, and streaming capabilities for large files. A URI‑mapping layer binds each URI to both a JCR node and a set of RDF triples, ensuring that updates to metadata are automatically reflected in the content repository and vice versa.
The framework embraces existing standards to maximize interoperability. It natively supports the PROV‑O ontology for provenance, FOAF for social relationships, GeoSPARQL for spatial data, and DCTerms/OWL for domain‑specific vocabularies. By combining these ontologies, a user can model a statement such as “On 12 May 2023, satellite sensor A collected temperature data (file X), which was processed by workflow B to produce result file Y, located at latitude 45.0 N, longitude ‑120.0 W.” Such a composite fact can be queried with a single SPARQL expression, retrieving the full lineage, geographic context, and temporal window in one step.
Three real‑world case studies illustrate Tupelo’s capabilities. In an astronomical data pipeline, distributed observatories upload raw images and associated observation metadata; Tupelo links these assets via provenance triples, enabling long‑term reproducibility and cross‑observatory discovery. In an environmental monitoring GIS, sensor streams of air‑quality measurements are ingested together with geolocation and timestamp information; analysts can query “all measurements for region X during the first half of 2023” and receive both the data files and the contextual metadata instantly. In a collaborative life‑science project, researchers share experimental data, analysis scripts, and manuscript drafts within a single Tupelo instance, applying role‑based access controls that restrict sensitive datasets to authorized personnel while still exposing provenance for auditability. Performance evaluations reported average SPARQL response times under 150 ms and stable throughput for files larger than 10 GB, with a 90 % reduction in metadata inconsistency errors compared to legacy file‑system approaches.
The authors acknowledge several challenges that remain. Scaling RDF triple stores to billions of statements will require advanced indexing and sharding strategies. Integrating heterogeneous domain ontologies can lead to semantic conflicts that must be resolved through ontology alignment tools. Long‑term digital preservation demands robust mechanisms for cryptographic signing and chain‑of‑trust verification, which are only partially addressed in the current prototype. Future work envisions extending Tupelo with distributed triple stores built on NoSQL platforms (e.g., Cassandra) and incorporating blockchain‑based integrity proofs to strengthen trust in archived scientific assets.
In conclusion, Tupelo demonstrates that a unified knowledge space—where global identifiers, RDF semantics, and an aggregatable content repository coexist—can effectively bridge the gap between data and its meaning in e‑Science environments. By providing a single, secure, and queryable platform for heterogeneous scientific assets, Tupelo enhances data discoverability, reproducibility, and collaborative potential, offering a scalable foundation for the next generation of data‑intensive research.
Comments & Academic Discussion
Loading comments...
Leave a Comment