Reconstructing the energy landscape of a distribution from Monte Carlo samples
Defining the energy function as the negative logarithm of the density, we explore the energy landscape of a distribution via the tree of sublevel sets of its energy. This tree represents the hierarchy among the connected components of the sublevel sets. We propose ways to annotate the tree so that it provides information on both topological and statistical aspects of the distribution, such as the local energy minima (local modes), their local domains and volumes, and the barriers between them. We develop a computational method to estimate the tree and reconstruct the energy landscape from Monte Carlo samples simulated at a wide energy range of a distribution. This method can be applied to any arbitrary distribution on a space with defined connectedness. We test the method on multimodal distributions and posterior distributions to show that our estimated trees are accurate compared to theoretical values. When used to perform Bayesian inference of DNA sequence segmentation, this approach reveals much more information than the standard approach based on marginal posterior distributions.
💡 Research Summary
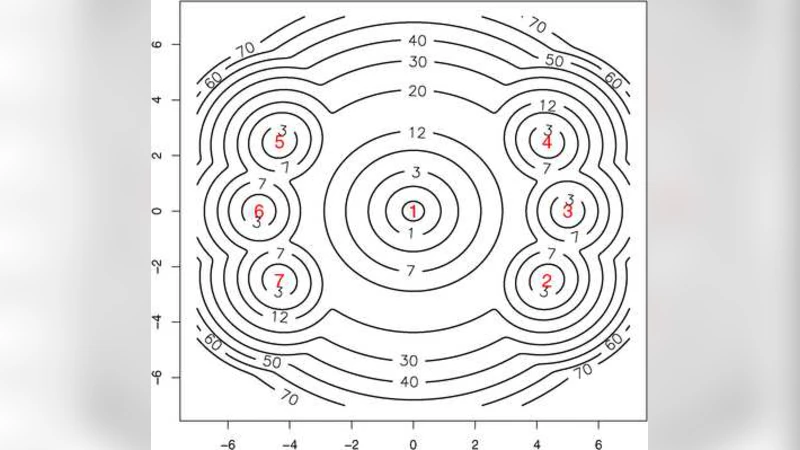
The paper introduces a novel framework for visualising and quantifying the structure of an arbitrary probability distribution by treating its negative log‑density as an energy function. The central construct is the “tree of sub‑level sets”: for each energy threshold ε, the set {x | E(x) ≤ ε} is examined, and its connected components are identified. As ε increases, smaller components merge into larger ones, and this merging process is recorded as a hierarchical tree. Each node of the tree corresponds to a connected component at a particular energy level, while parent‑child edges represent the moment when two components become connected as the energy threshold is raised. By annotating nodes with (i) the minimum energy (the local mode), (ii) the empirical volume (proportion of Monte‑Carlo samples belonging to the component), and (iii) the energy gap to the parent (the barrier height), the tree simultaneously encodes topological information (how many basins, how they are nested) and statistical information (how much probability mass each basin carries and how difficult it is to move between basins).
To estimate the tree from data, the authors first generate a large, diverse set of samples using advanced Monte‑Carlo schemes such as parallel tempering or temperature‑scaled Metropolis‑Hastings, ensuring coverage of both low‑ and high‑energy regions. The samples are sorted by their energy values. A graph is then built by connecting samples that are close in the underlying space, using either a k‑nearest‑neighbour rule or an ε‑ball neighbourhood. The Union‑Find (disjoint‑set) data structure is employed to maintain the current collection of connected components while the energy threshold is swept upward. Whenever two components become linked, the algorithm records the energy difference as the barrier height and creates a parent‑child relationship in the tree. The overall computational cost is dominated by the O(N log N) sorting step and the O(N α(N)) Union‑Find operations, where α is the inverse Ackermann function; the nearest‑neighbour search can be accelerated with approximate methods (e.g., locality‑sensitive hashing) for high‑dimensional data.
The authors validate the method on synthetic multimodal distributions where the exact energy landscape is analytically known. In two‑dimensional Gaussian mixture examples, the estimated tree reproduces the correct number of modes, the relative volumes of each basin, and the exact barrier heights, provided the sample size is sufficient. They then apply the technique to Bayesian inference problems, most notably DNA sequence segmentation. Traditional Bayesian segmentation reports marginal posterior probabilities for each position, which reveal where change‑points are likely but give little insight into the global segmentation structure. In contrast, the sub‑level set tree groups contiguous change‑points into basins, quantifies the total posterior mass of each segmentation hypothesis, and measures the energetic cost of switching between competing segmentations. This richer representation uncovers alternative segmentations that are statistically significant yet invisible to marginal analyses.
Beyond the case studies, the paper discusses several theoretical and practical implications. The tree offers a compact summary of the energy landscape that can be used for mode‑finding, for designing informed proposal distributions in Markov chain Monte Carlo, or for assessing convergence by monitoring the stability of the tree across iterations. It also provides a natural way to define “basin‑wise” credible regions, which are more informative than pointwise credible intervals in multimodal settings. Limitations include the difficulty of defining connectivity in very high dimensions, where the curse of dimensionality can render nearest‑neighbour graphs sparse and noisy, and the computational burden of generating enough samples to populate low‑probability basins. The authors suggest future work on dimensionality‑reduction techniques, adaptive neighbourhood constructions, and extensions to time‑varying energy landscapes in dynamical systems.
In summary, the paper contributes (1) a principled definition of the energy‑sublevel‑set tree as a unified topological‑statistical descriptor of any distribution, (2) an efficient algorithm for constructing this tree from Monte‑Carlo samples, and (3) compelling demonstrations that the tree reveals structural information—such as the number of modes, basin volumes, and inter‑mode barriers—that is inaccessible to conventional marginal or point‑estimate methods. This framework opens new avenues for Bayesian model exploration, multimodal optimisation, and the qualitative analysis of complex probabilistic models.
Comments & Academic Discussion
Loading comments...
Leave a Comment