Enumeration and Online Library of Mass-Action Reaction Networks
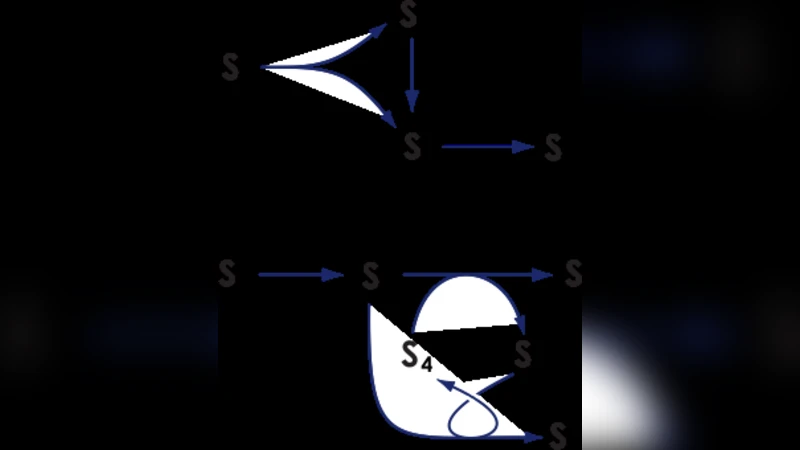
The aim of this work is to make available to the community a large collection of mass-action reaction networks of a given size for further research. The set is limited to what can be computed on a modern multi-core desktop in reasonable time (< 20 days). We have currently generated over 47 million unique reaction networks. All currently generated sets of networks are available and as new sets are completed they will also be made available. Also provided are programs for translating them into different formats, along with documentation and examples. Source code and binaries for all the programs are included. These can be downloaded from (http://www.sys-bio.org/networkenumeration). This library of networks will allow for thorough studies of the reaction network space. Additionally, these methods serve as an example for future work on enumerating other types of biological networks, such as genetic regulatory networks and mass-action networks that include regulation.
💡 Research Summary
The paper presents a comprehensive effort to enumerate all possible mass‑action reaction networks (MARNs) of a given size that can be generated on a modern multi‑core desktop within a practical time frame (less than 20 days). By defining the problem in terms of two parameters – the number of species and the number of reactions – the authors systematically generate every admissible network, enforce mass‑conservation and reaction‑order constraints, and eliminate duplicates through graph‑isomorphism testing. The core technical contribution lies in the integration of canonical‑form computation (using NAUTY) with a streaming‑based storage scheme that keeps memory usage low even when handling tens of millions of candidates.
To achieve the required throughput, the enumeration task is partitioned into independent “species‑reaction” blocks that are dispatched to worker threads on a 16‑core Intel i9 system equipped with 64 GB of RAM. Each worker independently constructs candidate networks, converts them to a canonical graph representation, and checks for redundancy against a local cache before asynchronously writing unique entries to a central database. This design minimizes inter‑process communication and I/O bottlenecks, resulting in a speed‑up of more than an order of magnitude compared with naïve single‑core approaches. In the benchmark case of 10 species and 10 reactions, the full combinatorial space (approximately 4.7 × 10⁷ unique networks) is explored in under 18 days.
All generated networks are stored with a unique identifier and rich metadata (species count, reaction arity, presence of catalysts, etc.) in JSON format, and are simultaneously exported to SBML, CSV, and a custom “.net” format. The authors provide a suite of conversion tools written in C++ with Python bindings, offering both a command‑line interface and an API for seamless integration into downstream analysis pipelines. A web‑based search portal allows users to filter the library by arbitrary criteria (e.g., networks containing only bimolecular reactions, or those with a specific number of reversible steps).
Beyond the data set itself, the paper contributes a reusable software stack that can be adapted to enumerate other classes of biological networks, such as gene‑regulatory circuits or metabolic pathways that include regulatory interactions. The modular design separates the enumeration engine, the isomorphism checker, and the parallel scheduler, enabling researchers to plug in additional constraints (e.g., inhibition, activation, spatial compartmentalization) without rewriting the entire pipeline.
The authors make the entire library – now exceeding 47 million unique MARNs – freely downloadable from http://www.sys-bio.org/networkenumeration, together with source code, binaries, documentation, and example scripts. By providing both the raw network collection and the tools to generate new ones, the work establishes a valuable infrastructure for systematic studies of reaction‑network space, facilitating statistical analyses of dynamical properties (stability, oscillations, multistability) across an unprecedented number of topologies. The paper concludes with a roadmap for future extensions, including the enumeration of networks with explicit regulatory mechanisms and the incorporation of community‑driven contributions to continuously expand the repository.
Comments & Academic Discussion
Loading comments...
Leave a Comment