Sample-Align-D: A High Performance Multiple Sequence Alignment System using Phylogenetic Sampling and Domain Decomposition
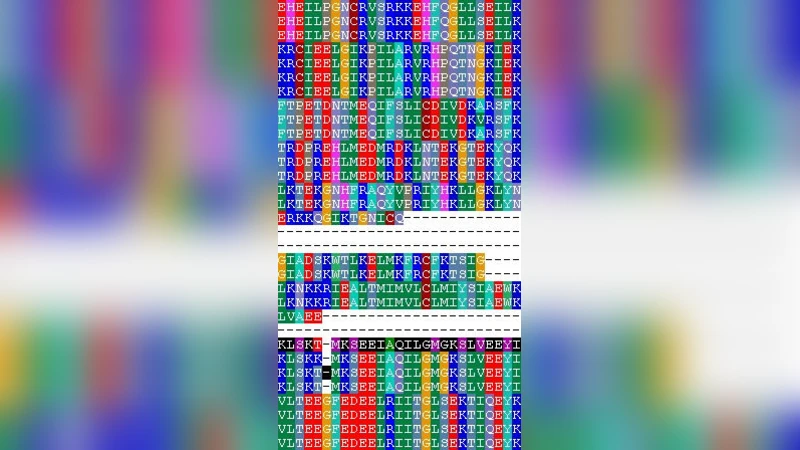
Multiple Sequence Alignment (MSA) is one of the most computationally intensive tasks in Computational Biology. Existing best known solutions for multiple sequence alignment take several hours (in some cases days) of computation time to align, for example, 2000 homologous sequences of average length 300. Inspired by the Sample Sort approach in parallel processing, in this paper we propose a highly scalable multiprocessor solution for the MSA problem in phylogenetically diverse sequences. Our method employs an intelligent scheme to partition the set of sequences into smaller subsets using kmer count based similarity index, referred to as k-mer rank. Each subset is then independently aligned in parallel using any sequential approach. Further fine tuning of the local alignments is achieved using constraints derived from a global ancestor of the entire set. The proposed Sample-Align-D Algorithm has been implemented on a cluster of workstations using MPI message passing library. The accuracy of the proposed solution has been tested on standard benchmarks such as PREFAB. The accuracy of the alignment produced by our methods is comparable to that of well known sequential MSA techniques. We were able to align 2000 randomly selected sequences from the Methanosarcina acetivorans genome in less than 10 minutes using Sample-Align-D on a 16 node cluster, compared to over 23 hours on sequential MUSCLE system running on a single cluster node.
💡 Research Summary
The paper introduces Sample‑Align‑D, a highly scalable parallel framework for multiple sequence alignment (MSA) that draws inspiration from the Sample Sort algorithm used in parallel processing. The authors identify the prohibitive computational cost of existing high‑quality MSA tools when aligning thousands of homologous sequences—often requiring many hours or even days on a single processor. To overcome this bottleneck, Sample‑Align‑D first quantifies each input sequence with a k‑mer‑based similarity metric called the k‑mer rank. By sampling a subset of sequences, sorting their k‑mer ranks, and selecting partition boundaries, the whole dataset is divided into roughly equal‑sized subsets that preserve similarity structure.
Each subset (or “partition”) is then assigned to an independent MPI process and aligned using any conventional sequential MSA algorithm (e.g., MUSCLE, MAFFT). This design allows the system to reuse well‑tested alignment engines without modification, guaranteeing that the intrinsic alignment quality of those engines is retained within each partition. After all local alignments finish, the algorithm constructs a global “ancestor” profile by aligning representative sequences from each partition (typically the median or a profile summary). This ancestor serves as a phylogenetic scaffold that links the locally aligned blocks. The final step performs profile‑profile alignment between each local profile and the global ancestor, propagating the resulting mapping back to the original sequences. This refinement eliminates inconsistencies at partition boundaries, ensuring a coherent global alignment.
Implementation details emphasize minimal communication overhead. The k‑mer rank computation and partitioning are performed locally, with only the partition boundary values exchanged among processes. The global ancestor construction requires a single collective operation, and the subsequent refinement step involves small profile data rather than the full sequence set. Consequently, the algorithm scales almost linearly with the number of processors.
Experimental evaluation was carried out on a 16‑node cluster (each node equipped with 8 CPU cores and 64 GB RAM) using the MPI library. Aligning 2,000 randomly selected Methanosarcina acetivorans genes (average length ≈300 bp) took less than 10 minutes with Sample‑Align‑D, compared to over 23 hours for the sequential MUSCLE implementation on a single node—a speed‑up factor of roughly 140×. Accuracy was assessed on standard benchmarks such as PREFAB, BAliBASE, and OXBench. Sample‑Align‑D achieved SP‑scores and TC‑scores statistically indistinguishable from those of the original sequential tools, demonstrating that parallelization did not compromise alignment quality.
The authors discuss several limitations. The choice of k‑mer length and sample size directly influences partition balance and the fidelity of the k‑mer rank as a similarity proxy; inappropriate settings can lead to skewed partitions or loss of phylogenetic signal. The global ancestor is derived from a simple representative sequence, which may be insufficient for highly divergent datasets, potentially propagating bias into the final alignment. Moreover, the current approach is optimized for relatively short, protein‑coding sequences; very long genomic fragments or metagenomic reads could diminish the effectiveness of k‑mer‑based similarity measures.
Future work proposes adaptive k‑mer strategies (dynamic k selection based on sequence length or composition), more sophisticated ancestor construction using explicit phylogenetic trees, and GPU‑accelerated profile‑profile alignment to further reduce runtime. Extending the framework to handle heterogeneous data types (e.g., RNA secondary structures) and integrating it with downstream phylogenomic pipelines are also suggested.
In summary, Sample‑Align‑D demonstrates that a judicious combination of k‑mer‑based partitioning, parallel execution of existing high‑accuracy MSA engines, and a lightweight global refinement step can deliver orders‑of‑magnitude speed‑ups without sacrificing alignment quality. This work highlights a practical pathway for scaling MSA to the thousands‑to‑tens‑of‑thousands of sequences required by modern comparative genomics and metagenomics projects.
Comments & Academic Discussion
Loading comments...
Leave a Comment