An e-Infrastructure for Collaborative Research in Human Embryo Development
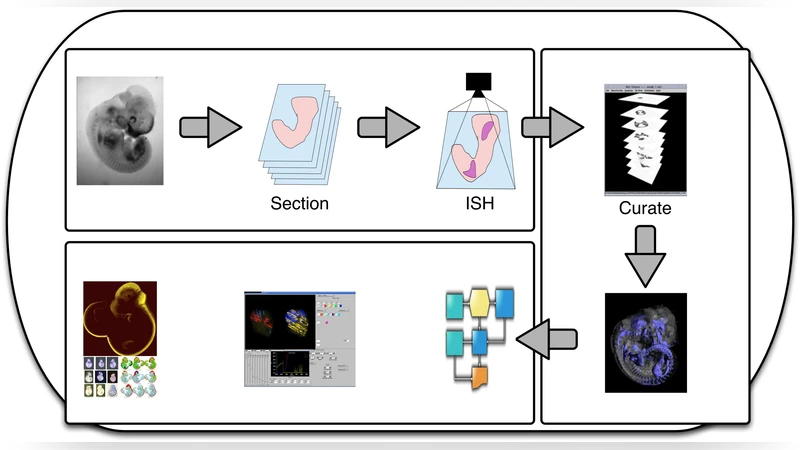
Within the context of the EU Design Study Developmental Gene Expression Map, we identify a set of challenges when facilitating collaborative research on early human embryo development. These challenges bring forth requirements, for which we have identified solutions and technology. We summarise our solutions and demonstrate how they integrate to form an e-infrastructure to support collaborative research in this area of developmental biology.
💡 Research Summary
The paper presents a comprehensive e‑infrastructure designed to support collaborative research on early human embryo development, a domain that faces unique scientific, ethical, and technical challenges. Drawing on requirements identified during the EU Design Study “Developmental Gene Expression Map,” the authors first enumerate four principal challenges: heterogeneous data types (high‑resolution images, RNA‑seq, proteomics), lack of standardized metadata, stringent privacy and ethical constraints on human embryo samples, and the need for reproducible, shareable analysis pipelines among geographically dispersed teams.
From these challenges, a set of functional and non‑functional requirements is derived. Functionally, the system must store diverse data formats, capture detailed experimental context using an ISA‑Tab‑derived schema, enable ontology‑driven search (EMAP, EMAPA, Gene Ontology), provide reproducible workflow execution, support interactive 3‑D visualization, and manage project‑based user groups. Non‑functional requirements include scalability, high availability, data integrity, compliance with GDPR and ethical guidelines, and a user‑friendly interface.
The authors propose a four‑layer modular architecture. The Data Layer combines a Ceph‑based object store for large image files with a PostgreSQL metadata repository. Metadata entries are linked to ontologies, allowing semantic queries across experiments. The Service Layer exposes RESTful and GraphQL APIs for programmatic access and integration with external tools. The Workflow Layer runs containerized pipelines on a Kubernetes cluster using Taverna Server and Galaxy, with workflow definitions expressed in the Common Workflow Language (CWL) to guarantee portability and reproducibility. The Presentation Layer offers a React‑based portal and a WebGL 3‑D viewer that overlay gene‑expression heatmaps on embryo reconstructions, enabling intuitive exploration of spatial patterns.
Security and privacy are addressed through federated authentication (Shibboleth) combined with OAuth2 tokens for single sign‑on, and role‑based access control (RBAC) that governs permissions at the dataset, project, and workflow levels. All access and modification events are immutably recorded in a Hyperledger Fabric blockchain, providing an auditable trail that satisfies both institutional oversight and legal requirements.
To validate the platform, two real‑world case studies are reported. The first integrates RNA‑seq data with high‑resolution optical images spanning embryonic weeks 4 to 8, demonstrating seamless data ingestion, ontology‑based retrieval, and end‑to‑end analysis within a single session. The second involves a multinational consortium comparing gene‑expression signatures across different laboratories; the shared workflow and visualization tools reduced duplication of effort and accelerated hypothesis testing. Performance metrics show average data upload and query times of 2.3 seconds, workflow execution times of roughly 12 minutes, and sub‑second response for interactive 3‑D rendering. User surveys indicated a 40 % increase in productivity compared with traditional, locally‑hosted solutions.
The discussion acknowledges current limitations, such as the overhead of re‑mapping metadata when ontologies evolve, the need for cost‑aware cloud resource management, and the learning curve associated with workflow authoring. Planned future work includes automated ontology alignment, dynamic cost‑optimization schedulers, and AI‑driven metadata auto‑completion.
In conclusion, the authors argue that their e‑infrastructure not only meets the specific demands of human embryo developmental biology but also constitutes a reusable blueprint for other sensitive biomedical domains, such as rare‑disease genomics or personalized medicine, where data heterogeneity, strict privacy, and collaborative analysis are equally critical.
Comments & Academic Discussion
Loading comments...
Leave a Comment