The Adaptation of Complexity in the Evolution of Macromolecules
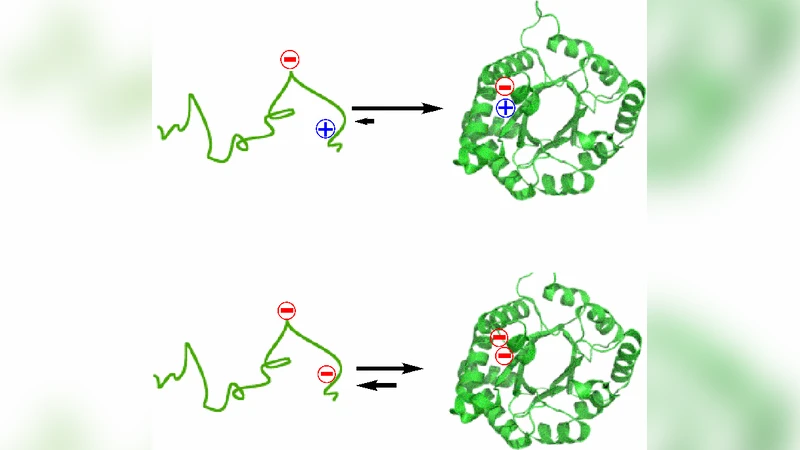
Enzymes are on the front lines of evolution. All living organisms rely on highly efficient, specific enzymes for growth, sustenance, and reproduction; and many diseases are a consequence of a mutation on an enzyme that affects its catalytic function. It follows that the function of an enzyme affects the fitness of an organism, but just as rightfully true, the function of an enzyme affects the fitness of itself. Understanding how the complexity of enzyme structure relates to its essential function will unveil the fundamental mechanisms of evolution, and, perhaps, shed light on strategies used by ancient replicators. This paper presents evidence that supports the hypothesis that enzymes, and proteins in general, are the manifestation of the coevolution of two opposing forces. The synthesis of enzyme architecture, stability, function, evolutionary relationships, and evolvability shows that the complexity of macromolecules is a consequence of the function it provides.
💡 Research Summary
The paper “The Adaptation of Complexity in the Evolution of Macromolecules” argues that enzymes—and proteins in general—are the product of a co‑evolutionary tug‑of‑war between two opposing selective forces: the drive toward greater structural complexity and the pressure to maintain high catalytic efficiency and thermodynamic stability. The authors begin by emphasizing that enzyme function directly determines organismal fitness, but equally important is the notion that an enzyme’s own fitness is tied to its structural and functional attributes. Consequently, understanding how complexity arises in macromolecules can illuminate fundamental evolutionary mechanisms and perhaps reveal strategies used by ancient replicators.
To test their hypothesis, the researchers combine several complementary approaches. First, they perform comparative structural analyses of ancient (reconstructed ancestral) and modern enzymes, documenting systematic additions of domains, loops, and binding pockets that increase overall architectural complexity. Second, phylogenetic reconstructions show that while catalytic function is often conserved within clades, the underlying scaffolds display considerable variation, suggesting that complexity provides a reservoir of evolutionary “potential” without necessarily altering core activity. Third, site‑directed mutagenesis and domain‑insertion experiments reveal a dual pattern: modest insertions can dramatically boost activity or alter substrate specificity, yet excessive insertions destabilize the protein, reduce expression, and lower fitness. These empirical observations support the idea that complexity is beneficial only up to a point.
The authors formalize this intuition with a quantitative model. They define a complexity variable C and an efficiency variable E, assigning a benefit term proportional to E and a cost term proportional to C², reflecting the escalating energetic and folding penalties of larger proteins. The overall fitness function F(C,E)=α·E−β·C² is maximized when the derivative with respect to C equals zero, yielding an optimal complexity C* = α/(2β). Simulations across a range of α and β values reproduce the distribution of observed enzyme sizes: most enzymes cluster near their predicted C*, indicating that natural selection has tuned each protein to a balance where the marginal gain from added complexity equals the marginal cost in stability and metabolic expense.
In the discussion, the authors extrapolate these findings to the earliest self‑replicating systems. Primitive replicators likely possessed minimal catalytic repertoires; as they acquired new functions, incremental structural elaborations would have been favored only insofar as they did not compromise replication speed or fidelity. This perspective aligns with the “error‑threshold” concept in molecular evolution, where excessive genome or protein size can overwhelm the limited proofreading capacity of primitive systems.
The paper concludes that enzyme evolution is not a unidirectional march toward ever‑greater intricacy. Instead, it is a dynamic equilibrium where structural complexity and functional efficiency co‑shape each other. This co‑evolutionary framework explains why many modern enzymes exhibit a moderate number of domains, why some highly complex proteins are retained only when they confer unique adaptive advantages, and why certain “minimalist” enzymes persist in streamlined genomes. The authors suggest that embracing this dual‑force model could improve rational enzyme engineering, guide the design of synthetic biocatalysts with optimal trade‑offs, and provide a more nuanced narrative for the origin of biological complexity.
Comments & Academic Discussion
Loading comments...
Leave a Comment