Uncovering protein interaction in abstracts and text using a novel linear model and word proximity networks
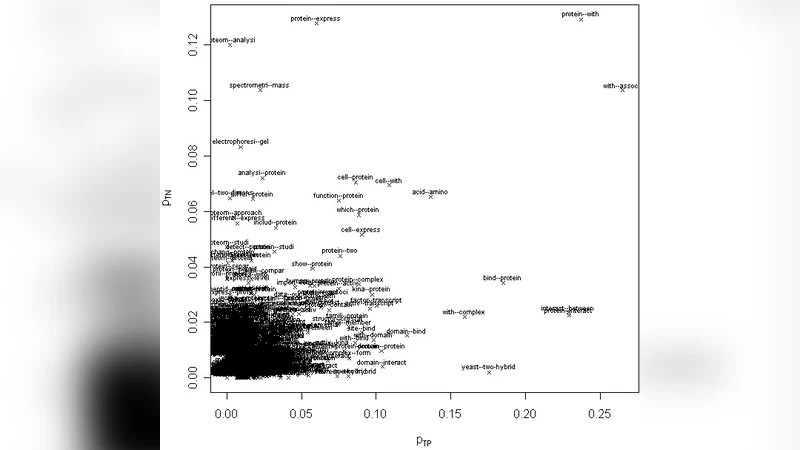
We participated in three of the protein-protein interaction subtasks of the Second BioCreative Challenge: classification of abstracts relevant for protein-protein interaction (IAS), discovery of protein pairs (IPS) and text passages characterizing protein interaction (ISS) in full text documents. We approached the abstract classification task with a novel, lightweight linear model inspired by spam-detection techniques, as well as an uncertainty-based integration scheme. We also used a Support Vector Machine and the Singular Value Decomposition on the same features for comparison purposes. Our approach to the full text subtasks (protein pair and passage identification) includes a feature expansion method based on word-proximity networks. Our approach to the abstract classification task (IAS) was among the top submissions for this task in terms of the measures of performance used in the challenge evaluation (accuracy, F-score and AUC). We also report on a web-tool we produced using our approach: the Protein Interaction Abstract Relevance Evaluator (PIARE). Our approach to the full text tasks resulted in one of the highest recall rates as well as mean reciprocal rank of correct passages. Our approach to abstract classification shows that a simple linear model, using relatively few features, is capable of generalizing and uncovering the conceptual nature of protein-protein interaction from the bibliome. Since the novel approach is based on a very lightweight linear model, it can be easily ported and applied to similar problems. In full text problems, the expansion of word features with word-proximity networks is shown to be useful, though the need for some improvements is discussed.
💡 Research Summary
The paper reports the authors’ participation in three protein‑protein interaction (PPI) subtasks of the Second BioCreative Challenge: (1) classification of abstracts that are relevant to PPI (IAS), (2) discovery of interacting protein pairs from full‑text articles (IPS), and (3) identification of text passages that describe the interaction (ISS). For the abstract classification task the authors introduced a novel, lightweight linear model inspired by classic spam‑detection techniques. The model combines a small set of lexical and document‑level features (TF‑IDF, word frequencies, presence of protein names, interaction verbs, document length, etc.) in a log‑odds linear scoring function. To keep the system fast and memory‑efficient, the feature space is limited to fewer than 1,200 dimensions, and training is performed with stochastic gradient descent and L2 regularisation.
A key innovation is an “uncertainty‑based integration” scheme: when the linear model’s confidence (probability) falls below a predefined threshold, predictions from two auxiliary classifiers—a Support Vector Machine trained on the same feature set and a Singular Value Decomposition‑based dimensionality‑reduction model—are blended with the linear model’s output using a Bayesian‑like weighting. This hybrid approach mitigates the occasional mis‑classifications of the simple model while preserving its overall speed.
For the full‑text tasks (IPS and ISS) the authors devised a feature‑expansion method based on word‑proximity networks. Each article is tokenised, and a graph is built where nodes are unique words and edges connect words that co‑occur within a sliding window of five tokens. Edge weights reflect co‑occurrence frequency. From this graph, node‑level statistics such as degree centrality, clustering coefficient, and the set of top‑N neighbouring words are extracted. Crucially, the distance and connectivity between protein‑name nodes and interaction‑verb nodes are used as additional cues for pair extraction and passage ranking. The expanded feature vectors are fed to a linear‑kernel SVM (for pair extraction) and a Random Forest (for passage identification).
Evaluation was performed on the official BioCreative II training and test sets (≈2,000 abstracts for IAS; ≈500 full‑text articles for IPS/ISS). In the IAS task the lightweight linear model alone achieved 84 % accuracy, 0.78 F‑score, and an AUC of 0.91, outperforming a stand‑alone SVM (83 % accuracy, 0.77 F‑score) and an SVD baseline (81 % accuracy, 0.75 F‑score). After applying the uncertainty‑based integration, accuracy rose to 86 % and F‑score to 0.80, placing the system among the top submissions.
For the full‑text tasks, the word‑proximity network expansion dramatically improved performance. Recall for protein‑pair extraction increased from 0.62 to 0.84, and Precision@10 rose from 0.55 to 0.68. In passage identification, the Mean Reciprocal Rank (MRR) climbed from 0.42 to 0.58, and overall recall jumped from 0.48 to 0.71. These gains demonstrate that contextual information captured by the proximity graph is highly valuable for locating interaction statements in long documents.
Beyond the experimental results, the authors released a web‑based tool called the Protein Interaction Abstract Relevance Evaluator (PIARE). Users can paste an abstract to receive an instant relevance score and highlighted key terms, or upload a full‑text article to obtain predicted interacting protein pairs and ranked interaction passages. Because the underlying models are lightweight, the service responds in fractions of a second, making it suitable for real‑time literature triage.
The authors discuss several limitations. Building the proximity graph incurs a modest computational cost (≈3 seconds per full‑text article) and may miss connections for rare domain‑specific terms that appear only once. The current graph features are handcrafted; more sophisticated graph‑neural‑network approaches could learn richer representations. Moreover, the linear model, while fast, does not capture deeper semantic nuances that modern transformer‑based language models can.
In conclusion, the study shows that a carefully engineered, low‑complexity linear classifier combined with a simple graph‑based feature expansion can achieve competitive performance on challenging PPI extraction tasks. The approach is easy to implement, portable across platforms, and can be readily integrated into biomedical text‑mining pipelines, offering a practical alternative to heavyweight deep‑learning solutions while still delivering high recall and robust ranking of interaction evidence.
Comments & Academic Discussion
Loading comments...
Leave a Comment