Self-organized periodicity of protein clusters in growing bacteria
Chemotaxis receptors in E. coli form clusters at the cell poles and also laterally along the cell body, and this clustering plays an important role in signal transduction. Recently, experiments using flourrescence imaging have shown that, during cell growth, lateral clusters form at positions approximately periodically spaced along the cell body. In this paper, we demonstrate within a lattice model that such spatial organization could arise spontaneously from a stochastic nucleation mechanism. The same mechanism may explain the recent observation of periodic aggregates of misfolded proteins in E. coli.
💡 Research Summary
The paper addresses a striking observation made by fluorescence microscopy: during the elongation of Escherichia coli cells, chemotaxis receptors and, more recently, aggregates of misfolded proteins appear not only at the poles but also laterally along the cell body, and these lateral clusters are spaced at roughly regular intervals (≈0.5–1 µm). The authors ask whether such spatial order can arise without any pre‑existing cellular scaffold or directed transport, i.e., purely from stochastic nucleation coupled to cell growth. To answer this, they construct a minimal lattice‑based stochastic model that captures three essential processes: (1) diffusion of individual protein monomers along a one‑dimensional lattice representing the longitudinal axis of the cell; (2) reversible binding of neighboring monomers to form small oligomers; and (3) a nucleation rule whereby an oligomer that reaches a critical size becomes a stable “nucleus” that dramatically lowers the dissociation rate of surrounding monomers, thereby acting as a seed for a larger cluster. The lattice length is increased at a constant rate to mimic exponential cell elongation, and new monomers are inserted uniformly to keep the average concentration constant.
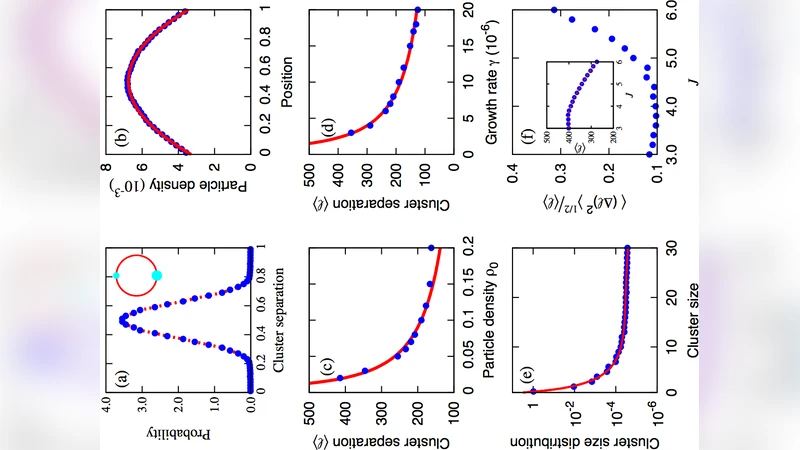
Simulations reveal a robust self‑organizing behavior. Initially a few nuclei appear randomly. As the lattice stretches, the distance between existing clusters grows, reducing the local monomer concentration near each nucleus and consequently lowering the nucleation probability in those regions. In the expanding gaps, the monomer concentration remains high enough that a new nucleus is likely to form. Once formed, this new cluster again partitions the cell into smaller segments, and the cycle repeats. The result is a quasi‑periodic arrangement of clusters whose average spacing depends on three tunable parameters: the critical nucleus size (or equivalently the nucleation free‑energy barrier), the monomer diffusion coefficient, and the cell‑elongation rate. By adjusting these parameters the model reproduces the experimentally observed spacing distribution and its modest variability.
The authors extend the same framework to aggregates of misfolded proteins, which are known to form “inclusion bodies” that also appear periodically in growing cells. By simply increasing the monomer production rate (to mimic stress‑induced over‑expression) while keeping the other parameters unchanged, the model generates periodic clusters of the same order of magnitude as those observed for the misfolded proteins. This suggests that the same physical mechanism—diffusion, stochastic nucleation, and growth‑driven dilution—may underlie both functional receptor clustering and pathological protein aggregation.
Key insights emerging from the study include: (i) a purely physical, stochastic nucleation process can generate spatial order without any cytoskeletal guidance or active positioning machinery; (ii) the interplay between cell elongation (which dilutes existing clusters) and nucleation kinetics naturally yields a characteristic length scale that sets the inter‑cluster distance; (iii) the model provides quantitative predictions for how changes in temperature, expression level, or mutation‑induced alterations in binding affinity would shift the spacing, offering a testable link between molecular biophysics and cellular morphology; and (iv) the same principle may be a universal strategy employed by bacteria to spatially segregate both signaling complexes and toxic aggregates, thereby optimizing signal transduction while minimizing interference from misfolded proteins.
In conclusion, the lattice model demonstrates that self‑organized periodicity of protein clusters in growing bacteria can emerge spontaneously from stochastic nucleation coupled to cell growth. The work bridges experimental observations with a simple yet powerful theoretical framework, opening avenues for experimental validation and for extending the concept to other cellular systems where periodic organization arises without explicit scaffolding.
Comments & Academic Discussion
Loading comments...
Leave a Comment