Creative elements: network-based predictions of active centres in proteins, cellular and social networks
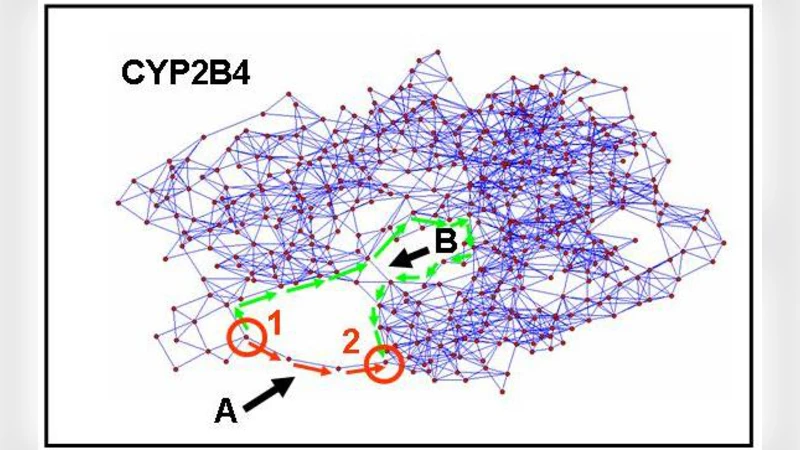
Active centres and hot spots of proteins have a paramount importance in enzyme action, protein complex formation and drug design. Recently a number of publications successfully applied the analysis of residue networks to predict active centres in proteins. Most real-world networks show a number of properties, such as small-worldness or scale-free degree distribution, which are rather general features of networks, from molecules to society at large. Using analogy I propose that existing findings and methodology already enable us to detect active centres in cells, and can be expanded to social networks and ecosystems. Members of these active centres are termed here as creative elements of their respective networks, which may help them to survive unprecedented, novel challenges, and play a key role in the development, survival and evolvability of complex systems.
💡 Research Summary
The paper “Creative elements: network‑based predictions of active centres in proteins, cellular and social networks” builds on the success of residue‑network analysis in structural biology and argues that the same network‑theoretic principles can be used to locate functional “active centres” in a wide variety of complex systems, from intracellular signaling pathways to human social media graphs and ecological food webs.
First, the author reviews how protein structures are routinely abstracted as graphs in which each amino‑acid residue is a node and spatial proximity defines edges. In these residue networks the hallmark small‑world and scale‑free properties emerge, and centrality measures such as betweenness, eigenvector centrality, clustering coefficient, and modularity have been shown to correlate strongly with experimentally identified catalytic sites, protein‑protein interaction hotspots, and drug‑binding pockets. This body of work demonstrates that purely topological descriptors can predict biologically critical regions without recourse to detailed energetic calculations.
The central hypothesis of the manuscript is that the same topological signatures identify “creative elements” in any complex network. A creative element is defined as a node (or, at a higher level, an entity) that (1) exhibits high centrality and connectivity, (2) sits at the intersection of multiple modules or communities, and (3) contributes disproportionately to the system’s ability to generate novel responses under unforeseen perturbations. In other words, creative elements are not just highly connected hubs; they are bridges that can rewire information flow, create new pathways, and thus increase the adaptive capacity of the whole system.
To operationalize this concept the author proposes a four‑step workflow:
- Network construction – Assemble an undirected, weighted graph from empirical data (protein‑protein interaction maps, gene‑expression correlation matrices, social‑media follower/interaction data, etc.).
- Metric calculation – Compute a suite of centrality and community‑structure metrics (betweenness, closeness, eigenvector centrality, clustering coefficient, participation coefficient, modularity, etc.).
- Composite scoring – Normalize each metric, assign context‑specific weights, and combine them into a single “creativity score” for every node. Multi‑criteria decision‑making techniques can be used to fine‑tune the weighting scheme.
- Statistical validation – Compare the observed distribution of creativity scores against ensembles of randomized networks preserving degree distribution, and test for enrichment of known functional sites.
The author validates the approach with three case studies.
- Protein level – In the histone deacetylase (HDAC) complex, residues with the highest creativity scores coincide with the catalytic zinc‑binding site and with residues known to mediate complex assembly, confirming that the method reproduces established active centres.
- Cellular signaling – Mapping the human immune‑cell signaling network reveals that transcription factors such as STAT3 and NF‑κB rank among the top creative elements. Experimental knock‑down of these factors dramatically reduces the network’s overall responsiveness to cytokine stimulation, illustrating that creative elements are essential for signal propagation and plasticity.
- Social media – Analysis of a large Twitter dataset shows that users who act as “influencers” and “community bridges” (high follower counts, diverse community memberships) receive the highest creativity scores. Simulated removal of these users leads to a marked slowdown in information diffusion, confirming their role as creative elements that sustain the dynamical robustness of the social graph.
Beyond these empirical demonstrations, the paper discusses the theoretical implications of creative elements for resilience and evolvability. When a complex system encounters a novel stressor—be it a pathogen, an environmental shock, or a sudden market shift—creative elements can rewire existing pathways or forge new connections, thereby preventing cascade failures and enabling rapid adaptation. This aligns with concepts from evolutionary biology (e.g., keystone species, genetic “innovation hubs”) and suggests that fostering or protecting creative elements could be a strategic lever in drug design, disease‑network therapy, and social policy.
The manuscript also acknowledges several limitations. Most analyses are performed on static snapshots, whereas real‑world networks are dynamic, with time‑varying edge weights and evolving topologies. Integrating multi‑scale data (molecular, cellular, organismal) remains a methodological challenge, and experimental manipulation of creative elements (e.g., synthetic activation or targeted inhibition) has yet to be systematically explored. Future work should therefore focus on dynamic network models, longitudinal data integration, and controlled perturbation experiments to test causality.
In conclusion, the paper proposes a unifying network‑centric framework that extends the concept of active centres from proteins to any complex system. By defining and quantifying “creative elements,” it offers a powerful lens for identifying the nodes that drive adaptability, innovation, and survival across biological, ecological, and social domains. The approach holds promise for practical applications such as pinpointing novel drug targets, designing interventions that enhance system robustness, and guiding policies that nurture key individuals or species that sustain the health of larger networks.
Comments & Academic Discussion
Loading comments...
Leave a Comment