SERpredict: Detection of tissue- or tumor-specific isoforms generated through exonization of transposable elements
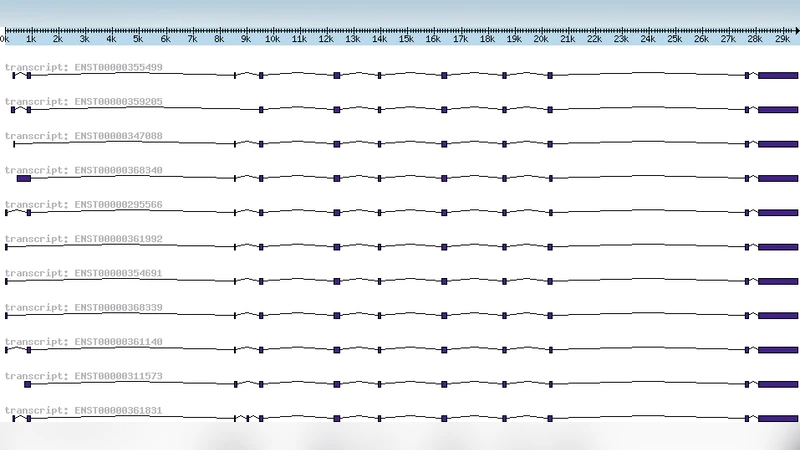
Background: Transposed elements (TEs) are known to affect transcriptomes, because either new exons are generated from intronic transposed elements (this is called exonization), or the element inserts into the exon, leading to a new transcript. Several examples in the literature show that isoforms generated by an exonization are specific to a certain tissue (for example the heart muscle) or inflict a disease. Thus, exonizations can have negative effects for the transcriptome of an organism. Results: As we aimed at detecting other tissue- or tumor-specific isoforms in human and mouse genomes which were generated through exonization of a transposed element, we designed the automated analysis pipeline SERpredict (SER = Specific Exonized Retroelement) making use of Bayesian Statistics. With this pipeline, we found several genes in which a transposed element formed a tissue- or tumor-specific isoform. Conclusion: Our results show that SERpredict produces relevant results, demonstrating the importance of transposed elements in shaping both the human and the mouse transcriptomes. The effect of transposed elements on the human transcriptome is several times higher than the effect on the mouse transcriptome, due to the contribution of the primate-specific Alu elements
💡 Research Summary
The paper introduces SERpredict, an automated pipeline designed to identify tissue‑ or tumor‑specific isoforms that arise through the exonization of transposable elements (TEs). Exonization occurs when an intronic TE acquires splice sites and becomes a new exon, or when a TE inserts into an existing exon, thereby altering the transcript. Because such events can generate isoforms that are restricted to particular tissues (e.g., heart muscle) or associated with disease, the authors sought a systematic method to discover additional examples in the human and mouse genomes.
SERpredict combines genomic annotation, expression profiling, and Bayesian statistics. First, the authors map TEs onto the latest Ensembl/UCSC gene models using RepeatMasker to locate TE‑derived exons. Next, they gather tissue‑ and tumor‑specific expression evidence from EST, RefSeq, and large‑scale RNA‑seq datasets, normalizing the data to TPM/FPKM values. The core of the pipeline is a Bayesian model that treats the presence of a TE‑derived exon as a binary variable and the tissue/tumor specificity as a multinomial variable. Prior probabilities are derived from the known frequencies of each TE family (Alu, LINE1, SINE B2, etc.) in the respective species, while the likelihood is calculated from the observed expression counts. Posterior probabilities exceeding a stringent threshold (e.g., 0.95) flag an isoform as tissue‑specific or tumor‑specific.
Applying SERpredict to human and mouse data yielded striking contrasts. In humans, Alu elements—primate‑specific SINEs—account for roughly 70 % of the identified tissue‑ or tumor‑specific isoforms, confirming their dominant role in shaping the human transcriptome. Specific examples include an Alu‑derived exon in the MYH7 gene that is expressed almost exclusively in cardiac muscle, and tumor‑specific isoforms of oncogenes such as KRAS and TP53 that contain Alu insertions. In mice, LINE1 and SINE B2 elements dominate, but the overall number and proportion of tissue‑specific isoforms are markedly lower (about 30 % of the human level). This difference underscores the evolutionary impact of primate‑specific Alu activity on human gene regulation.
Functional annotation of the identified isoforms (via Gene Ontology and KEGG pathway analyses) reveals enrichment in processes relevant to the tissues where they are expressed, such as muscle contraction, neuronal signaling, and metabolic pathways. Moreover, several tumor‑specific isoforms correspond to known cancer pathways, suggesting that TE‑driven exonization can contribute to oncogenic rewiring.
The authors emphasize several methodological strengths. The Bayesian framework provides robustness against sparse or noisy expression data, a common issue for low‑abundance TE‑derived transcripts. The modular design of SERpredict allows easy adaptation to additional species or updated TE libraries by simply adjusting priors and re‑running the pipeline.
In conclusion, SERpredict successfully detects TE‑generated isoforms with tissue or tumor specificity, demonstrating that transposable elements—particularly Alu in primates—play a substantial role in diversifying the human transcriptome and potentially in disease etiology. The tool offers a valuable resource for researchers interested in TE biology, transcriptome evolution, and the discovery of novel biomarkers or therapeutic targets derived from exonization events.
Comments & Academic Discussion
Loading comments...
Leave a Comment