On the role of conserved moieties in shaping the robustness and production capabilities of reaction networks
We study a simplified, solvable model of a fully-connected metabolic network with constrained quenched disorder to mimic the conservation laws imposed by stoichiometry on chemical reactions. Within a spin-glass type of approach, we show that in presence of a conserved metabolic pool the flux state corresponding to maximal growth is stationary independently of the pool size. In addition, and at odds with the case of unconstrained networks, the volume of optimal flux configurations remains finite, indicating that the frustration imposed by stoichiometric constraints, while reducing growth capabilities, confers robustness and flexibility to the system. These results have a clear biological interpretation and provide a basic, fully analytical explanation to features recently observed in real metabolic networks.
💡 Research Summary
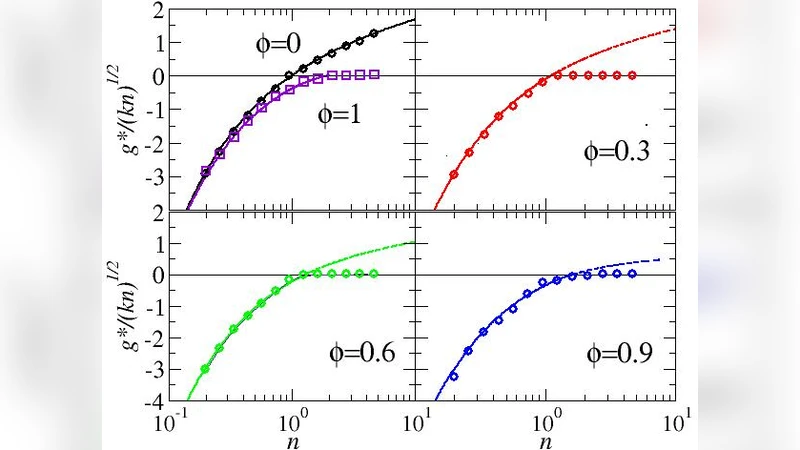
The paper presents a fully analytical study of a stylized, fully‑connected metabolic network in which the stoichiometric conservation laws are represented by quenched disorder constraints. By mapping the problem onto a spin‑glass–type statistical‑mechanical model and employing the replica method, the authors derive closed‑form expressions for the maximal growth rate and for the volume of flux configurations that achieve this optimum. A central ingredient of the model is a conserved metabolic pool (a set of metabolites whose total amount is fixed by stoichiometry). The size of this pool is denoted by the parameter ϕ, representing the fraction of the total metabolite mass that belongs to the conserved set.
The analysis yields two striking results. First, the optimal growth rate g_max is completely independent of ϕ. In other words, regardless of how large or small the conserved pool is, the direction of the flux vector that maximizes growth remains stationary. This reflects the fact that the conservation law eliminates certain degrees of freedom but does not alter the direction in flux space that yields the highest possible biomass production. Second, the space of optimal fluxes retains a finite volume when a conserved pool is present. In unconstrained random networks the optimal solution collapses to a single point (zero volume), indicating a unique flux configuration. By contrast, the presence of the conserved pool introduces frustration that prevents a unique solution, leaving a multidimensional manifold of equally optimal fluxes. Mathematically, the variance Δ of the optimal fluxes decreases with increasing ϕ but never vanishes, confirming that multiple optimal states survive.
Biologically, these findings provide a theoretical underpinning for observations made in real cellular metabolism. Real organisms possess several conserved pools – for example, the adenine nucleotide pool (ATP, ADP, AMP) or pools of certain amino acids – that are tightly regulated. The model shows that such pools inevitably reduce the maximal attainable growth (the “frustration” effect) but simultaneously endow the metabolic network with robustness: the system can accommodate perturbations, gene knock‑outs, or environmental fluctuations by moving within the finite manifold of optimal fluxes rather than being forced into a single brittle configuration. This dual effect explains why metabolic networks are both efficient and flexible, a hallmark of evolutionary adaptation.
The authors acknowledge the simplifications inherent in their approach. The assumption of full connectivity ignores the sparse, modular architecture of actual metabolic graphs, and the random assignment of reaction efficiencies does not capture enzyme kinetics or regulatory feedback. Nevertheless, the model’s tractability allows a clear isolation of the impact of stoichiometric conservation, something that would be difficult to achieve in more detailed simulations.
Future work suggested by the authors includes (i) embedding realistic network topologies derived from genome‑scale reconstructions, (ii) incorporating dynamic regulation (transcriptional control, allosteric inhibition) into the statistical‑mechanical framework, and (iii) comparing the analytical predictions with flux‑balance analysis and experimental ^13C‑labeling data. Such extensions would bridge the gap between the elegant, solvable model presented here and the complex, data‑rich reality of cellular metabolism, with potential applications in metabolic engineering (designing robust production strains) and in medicine (identifying metabolic vulnerabilities in disease states).
Comments & Academic Discussion
Loading comments...
Leave a Comment