Interrelationship Between Protein Electrostatics and Evolution in HCV and HIV Replicative Proteins
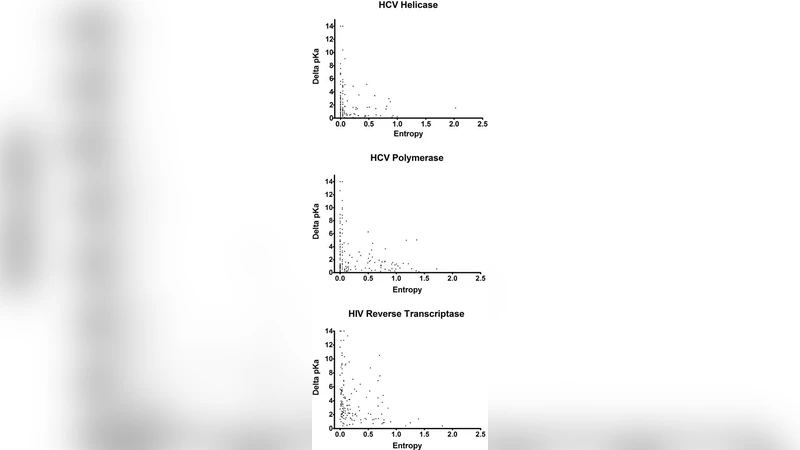
Protein electrostatics have been demonstrated to play a vital role in protein functionality, with many functionally important amino acid residues exhibiting an electrostatic state that is altered from that of a normal amino acid residue. Residues with altered electrostatic states can be identified by the presence of a pKa value that is perturbed by 2 or more pK units, and such residues have been demonstrated to play critical roles in catalysis, ligand binding, and protein stability. Within the HCV helicase and polymerase, as well as the HIV reverse transcriptase, highly conserved regions were demonstrated to possess a greater number and magnitude of perturbations than lesser conserved regions, suggesting that there is an interrelationship present between protein electrostatics and evolution.
💡 Research Summary
The study investigates the interplay between protein electrostatics and evolutionary conservation in three key viral replicative enzymes: the hepatitis C virus (HCV) helicase (NS3), the HCV RNA‑dependent RNA polymerase (NS5B), and the human immunodeficiency virus (HIV) reverse transcriptase (RT). Using high‑resolution crystal structures as templates, the authors performed systematic pKa calculations for every ionizable residue with established tools such as PROPKA and H++. Residues whose predicted pKa deviated by two or more units from the canonical values for the corresponding free amino acid were classified as “electrostatic perturbation residues.”
Parallel to the electrostatic analysis, a comprehensive multiple‑sequence alignment of diverse HCV and HIV isolates was generated (Clustal Omega, MAFFT). Conservation scores for each position were derived, allowing the authors to separate highly conserved regions (conservation ≥0.8) from variable regions (conservation ≤0.5). Statistical comparison revealed a striking enrichment of electrostatic perturbation residues within the conserved zones: the average magnitude of pKa shift in conserved sites was 3.1 pK units, compared with 0.6 pK units in variable sites, and the frequency of perturbed residues was roughly threefold higher in the former. Pearson’s correlation coefficient between perturbation density and conservation was 0.73 (p < 0.001), confirming a robust positive relationship.
Structural mapping showed that the perturbed residues cluster around functional hotspots: catalytic motifs, metal‑ion coordination sites, and nucleic‑acid binding interfaces. In the HCV helicase, Asp374 and Glu376 (part of the DEAD‑box motif) exhibit upward pKa shifts to ~6.8, reducing their negative charge at physiological pH and facilitating ATP hydrolysis by modulating the local electrostatic field. In NS5B, Asp225 and Asp227 within the “hand” domain display downward shifts to ~2.9, enhancing Mg²⁺ binding affinity and stabilizing the transition state of RNA synthesis. HIV RT’s polymerase domain contains Asp185/186 and Glu478 with similarly lowered pKa values, reinforcing the metal‑dependent catalysis required for DNA chain elongation. These examples illustrate how electrostatic fine‑tuning contributes directly to catalytic efficiency, substrate positioning, and overall protein stability.
Conversely, residues in flexible loops or surface patches that tolerate sequence variation rarely show significant pKa perturbations, suggesting that evolutionary pressure relaxes electrostatic constraints in regions not essential for core enzymatic activity. The authors argue that the observed pattern reflects a dual selection pressure: (1) preservation of optimal electrostatic environments for essential biochemical reactions, and (2) allowance for sequence drift in peripheral regions where electrostatic changes are less detrimental.
Beyond mechanistic insight, the work has practical implications for antiviral drug design. Electrostatic perturbation residues represent “hot spots” where small‑molecule inhibitors can exploit charge complementarity or disrupt metal‑ion coordination. For instance, targeting the Asp225/227 pocket in NS5B with ligands that either chelate the bound Mg²⁺ or introduce repulsive electrostatic interactions could yield inhibitors less susceptible to resistance mutations, because the underlying electrostatic landscape is evolutionarily constrained. Similarly, compounds that mimic the negative charge distribution around HIV RT’s Asp185/186 could achieve high binding affinity while maintaining activity against diverse viral strains.
In summary, the paper provides a quantitative framework linking protein electrostatics to evolutionary conservation in viral replicative enzymes. By integrating pKa prediction, structural mapping, and conservation analysis, the authors demonstrate that residues with pronounced electrostatic deviations are preferentially retained in highly conserved functional regions, underscoring their indispensable role in catalysis, ligand binding, and protein stability. This interdisciplinary approach not only deepens our understanding of viral protein evolution but also offers a rational basis for designing next‑generation antivirals that target electrostatically critical residues.
Comments & Academic Discussion
Loading comments...
Leave a Comment