Ribosome collisions and Translation efficiency: Optimization by codon usage and mRNA destabilization
Individual mRNAs are translated by multiple ribosomes that initiate translation with a few seconds interval. The ribosome speed is codon dependant, and ribosome queuing has been suggested to explain specific data for translation of some mRNAs in vivo. By modelling the stochastic translation process as a traffic problem, we here analyze conditions and consequences of collisions and queuing. The model allowed us to determine the on-rate (0.8 to 1.1 initiations per sec) and the time (1 sec) the preceding ribosome occludes initiation for Escherichia coli lacZ mRNA in vivo. We find that ribosome collisions and queues are inevitable consequences of a stochastic translation mechanism that reduce the translation efficiency substantially on natural mRNAs. The cells minimize collisions by having its mRNAs being unstable and by a highly selected codon usage in the start of the mRNA. The cost of mRNA breakdown is offset by the concomitant increase in translational efficiency.
💡 Research Summary
The paper presents a quantitative analysis of how ribosome collisions and mRNA instability jointly shape translation efficiency in bacteria, using the Escherichia coli lacZ messenger as a model system. The authors recast translation as a stochastic traffic problem, employing a total‑asymmetric simple exclusion process (TASEP) framework in which ribosomes are particles that enter the mRNA at an initiation rate (k_on), move forward with codon‑specific stepping rates (v_i), and occupy a finite footprint that blocks subsequent initiations for a defined occlusion time (τ_occl). By fitting the model to ribosome profiling (Ribo‑seq) data and measured mRNA half‑life, they estimate an in‑vivo initiation frequency of 0.8–1.1 initiations per second and an occlusion time of roughly one second, meaning that a ribosome shields the start codon for about 30 nucleotides after it has bound.
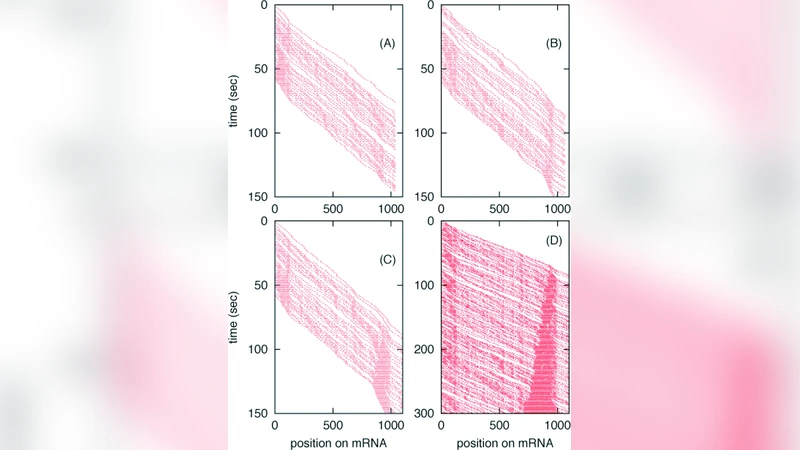
Simulation results reveal three key phenomena. First, because elongation speeds differ among codons, stochastic variations accumulate downstream, creating “traffic jams” where trailing ribosomes pause behind slower ones. These jams markedly reduce the overall protein output, with predicted efficiency losses of 20–30 % when slow codons appear early in the transcript. Second, the codon composition at the very beginning of the mRNA is a powerful lever: an enrichment of fast‑translating codons (e.g., GAA, GAG) minimizes early queuing and improves throughput, whereas the presence of slow codons near the start dramatically raises collision probability. Third, the authors demonstrate that a short mRNA half‑life—i.e., rapid degradation—lowers the average ribosome density on a transcript, thereby decreasing the chance of collisions. In effect, unstable mRNAs can paradoxically increase translational efficiency by limiting the window in which ribosome traffic can become congested.
The discussion interprets these findings in an evolutionary context. Highly expressed genes tend to combine a “fast‑start” codon bias with relatively brief mRNA lifespans, suggesting that natural selection has tuned both sequence composition and transcript stability to mitigate the energetic and fidelity costs associated with ribosome collisions. From a synthetic biology perspective, the work provides concrete design rules: (1) place optimal, fast‑translating codons in the first 30–50 nucleotides of any engineered open reading frame, and (2) modulate mRNA stability (for example, by adjusting 5′‑UTR elements or incorporating degradation tags) to keep ribosome density at a level that prevents queuing while still delivering sufficient protein yield.
Overall, the study advances our mechanistic understanding of translation by integrating kinetic modeling, experimental data, and evolutionary theory. It underscores that translation efficiency is not solely dictated by codon usage or ribosome availability, but also by the dynamic interplay between ribosome traffic and mRNA turnover. The presented model offers a versatile platform for predicting how changes in codon composition, initiation rates, or mRNA decay rates will impact protein synthesis, thereby informing both basic research on gene regulation and the rational design of high‑efficiency expression constructs.
Comments & Academic Discussion
Loading comments...
Leave a Comment