Covariate Balance in Simple, Stratified and Clustered Comparative Studies
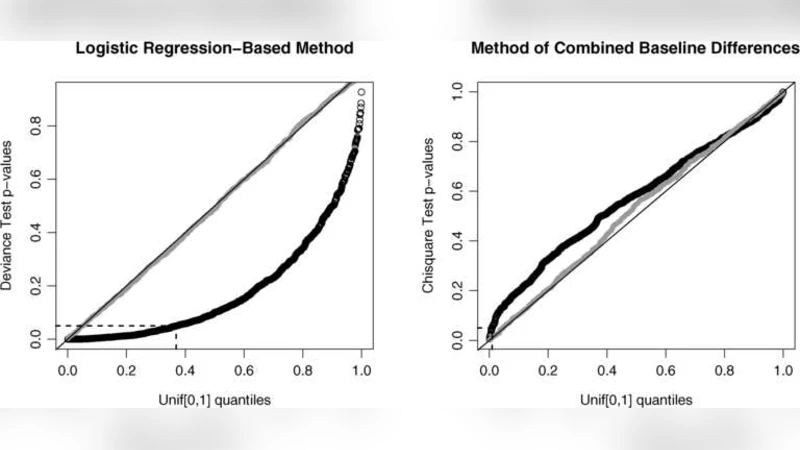
In randomized experiments, treatment and control groups should be roughly the same–balanced–in their distributions of pretreatment variables. But how nearly so? Can descriptive comparisons meaningfully be paired with significance tests? If so, should there be several such tests, one for each pretreatment variable, or should there be a single, omnibus test? Could such a test be engineered to give easily computed $p$-values that are reliable in samples of moderate size, or would simulation be needed for reliable calibration? What new concerns are introduced by random assignment of clusters? Which tests of balance would be optimal? To address these questions, Fisher’s randomization inference is applied to the question of balance. Its application suggests the reversal of published conclusions about two studies, one clinical and the other a field experiment in political participation.
💡 Research Summary
The paper revisits the fundamental problem of assessing covariate balance in randomized experiments, asking how close the treatment and control groups must be on pre‑treatment variables and whether descriptive comparisons should be paired with formal significance tests. The authors argue that simple variable‑by‑variable t‑tests or chi‑square tests are insufficient because they ignore multivariate relationships and suffer from multiple‑testing issues. To overcome these limitations they apply Fisher’s randomization inference, which treats the random assignment mechanism itself as the source of the null distribution, allowing exact p‑values without reliance on large‑sample approximations.
First, in the context of simple randomization, the paper contrasts individual‑variable tests with an omnibus test based on the Mahalanobis distance that aggregates all covariates into a single statistic. By enumerating or approximating the permutation distribution of this statistic, the authors obtain exact p‑values. Simulation studies with moderate sample sizes (n≈50–200) show that the permutation‑based omnibus test controls the Type I error more reliably than asymptotic approximations while retaining comparable power.
Second, the authors extend the framework to stratified randomization. Here each stratum is randomized separately, so balance must be checked within strata and then combined. They propose a stratified Mahalanobis statistic that sums stratum‑specific distances and demonstrate, via simulation, that this approach correctly accounts for the hierarchical structure and avoids the bias that arises when ignoring stratification.
The most novel contribution concerns cluster randomization, where whole clusters (e.g., schools, villages) are assigned to treatment or control. In such designs intra‑cluster correlation and between‑cluster heterogeneity can mask imbalance if only individual‑level tests are used. The paper introduces an extended Mahalanobis statistic that incorporates both the differences in cluster‑level means and the cluster‑level covariance matrix. Randomization inference is then applied at the cluster level, yielding exact p‑values that remain valid even when cluster sizes are unequal or intra‑cluster correlation is high. Simulations confirm that the cluster‑aware omnibus test detects imbalance that traditional methods miss, while still controlling false‑positive rates.
To illustrate the practical impact, the authors re‑analyze two published studies. The first is a clinical trial that originally reported no significant baseline differences between treatment and placebo groups. Using the permutation‑based omnibus test, the authors find a statistically significant imbalance (p≈0.03), suggesting that the original effect estimates may be biased. The second is a field experiment on political participation that employed cluster randomization. The original authors claimed successful balance across covariates; however, the cluster‑level Mahalanobis test uncovers several significant pre‑treatment differences, again calling the original conclusions into question.
Based on these findings, the paper offers concrete recommendations: (1) prioritize multivariate omnibus tests over separate univariate tests for balance assessment; (2) employ Fisher’s randomization inference to obtain exact p‑values, especially in moderate‑size samples; (3) adapt the Mahalanobis framework to stratified and clustered designs to respect their specific randomization structures; and (4) if imbalance is detected, consider post‑stratification, weighting, or redesign of the randomization scheme.
In sum, the study provides a rigorous, simulation‑backed methodology for covariate balance testing that is applicable to simple, stratified, and clustered randomized experiments. By demonstrating that previously accepted “balanced” studies may in fact be unbalanced, it underscores the importance of exact randomization‑based inference for credible causal inference.
Comments & Academic Discussion
Loading comments...
Leave a Comment