Conditional probability based significance tests for sequential patterns in multi-neuronal spike trains
In this paper we consider the problem of detecting statistically significant sequential patterns in multi-neuronal spike trains. These patterns are characterized by ordered sequences of spikes from different neurons with specific delays between spikes. We have previously proposed a data mining scheme to efficiently discover such patterns which are frequent in the sense that the count of non-overlapping occurrences of the pattern in the data stream is above a threshold. Here we propose a method to determine the statistical significance of these repeating patterns and to set the thresholds automatically. The novelty of our approach is that we use a compound null hypothesis that includes not only models of independent neurons but also models where neurons have weak dependencies. The strength of interaction among the neurons is represented in terms of certain pair-wise conditional probabilities. We specify our null hypothesis by putting an upper bound on all such conditional probabilities. We construct a probabilistic model that captures the counting process and use this to calculate the mean and variance of the count for any pattern. Using this we derive a test of significance for rejecting such a null hypothesis. This also allows us to rank-order different significant patterns. We illustrate the effectiveness of our approach using spike trains generated from a non-homogeneous Poisson model with embedded dependencies.
💡 Research Summary
The paper addresses the challenging problem of identifying statistically significant sequential firing patterns in multi‑neuronal spike trains. Such patterns consist of ordered spikes from distinct neurons separated by specific inter‑spike delays. While previous work by the authors introduced an efficient data‑mining algorithm that extracts patterns whose non‑overlapping occurrence count exceeds a user‑defined threshold, that approach lacked a principled statistical test for significance and relied on an overly simplistic null model assuming complete independence among neurons.
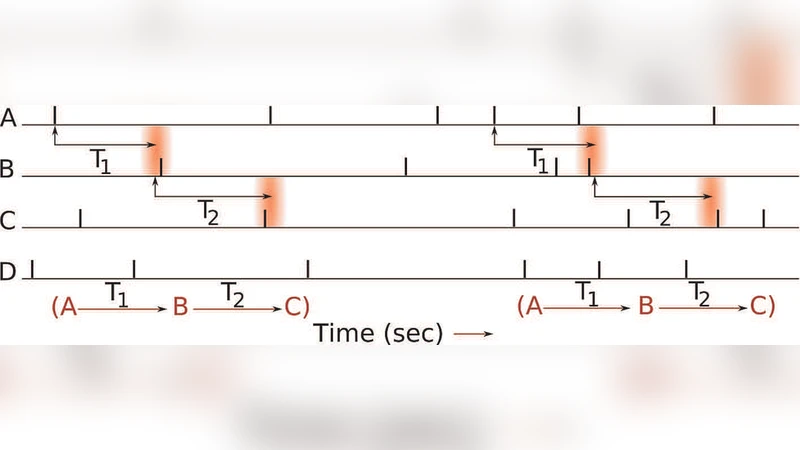
To overcome these limitations, the authors propose a novel hypothesis‑testing framework that (1) defines a compound null hypothesis encompassing not only independent firing but also weakly dependent neuronal interactions, and (2) derives an exact probabilistic model for the counting process of pattern occurrences. The key idea is to bound all pairwise conditional probabilities (P(j\mid i,\tau)) – the probability that neuron (j) fires (\tau) milliseconds after neuron (i) – by a common upper limit (\theta). When (\theta=0) the null hypothesis reduces to the classic independence model; larger values of (\theta) permit modest dependencies, thereby making the test robust to realistic, low‑level correlations that are ubiquitous in neural data.
Under this null, each potential start time of a pattern can be regarded as a Bernoulli trial whose success probability is the product of the relevant conditional probabilities. By treating the entire recording of length (T) as a sequence of such trials, the authors obtain closed‑form expressions for the expected count (\mu) and variance (\sigma^{2}) of any candidate pattern. The derivation uses a Markov‑chain representation of the sequential dependencies and accounts for the non‑overlapping constraint by effectively “resetting” the process after each successful detection. With (\mu) and (\sigma^{2}) in hand, a significance test is constructed: the observed count is compared to the theoretical distribution (approximated by a normal distribution for large (T) or by the exact binomial/Poisson distribution when appropriate). The resulting p‑value quantifies the probability of observing at least as many occurrences under the null hypothesis.
Because the null hypothesis is parameterized by (\theta), the method automatically yields an appropriate detection threshold for any desired level of permissible inter‑neuronal coupling. Moreover, the p‑values provide a natural ranking of patterns: those with the smallest p‑values are the most unlikely under the null and thus the most compelling candidates for genuine functional connectivity.
The authors validate their approach using synthetic spike trains generated from a non‑homogeneous Poisson process. They embed specific dependencies by setting selected conditional probabilities above the background level and then vary (\theta) and pattern length to assess detection power and false‑positive rates. Results demonstrate that (i) the proposed test maintains high power even when dependencies are weak, (ii) it dramatically reduces false positives compared with a naïve fixed‑count threshold, and (iii) the ranking of patterns correlates strongly with the ground‑truth strength of the implanted connections.
In the discussion, the authors acknowledge that the current formulation only captures pairwise conditional probabilities and does not directly model higher‑order interactions (e.g., triplet or quadruplet dependencies). They suggest extending the framework to a Bayesian setting where priors over higher‑order motifs could be incorporated, and they propose adapting the method for online, streaming analysis of neural data.
Overall, the paper makes a substantial contribution to neural data mining by introducing a rigorous statistical test that bridges efficient pattern discovery with principled significance assessment. By allowing a controlled amount of weak dependence in the null model, the method aligns more closely with the physiological reality of neuronal networks, offering researchers a powerful tool to uncover truly meaningful sequential firing motifs.
Comments & Academic Discussion
Loading comments...
Leave a Comment