Power-law scaling in protein synthesis of a stochastic regulon
We investigate the protein expression pattern of the lamB gene in Escherichia coli LE392. The gene product LamB is an important membrane protein for maltose transport into cells but it is also exploited by bacteriophage lambda for infection. Although our bacterial population is clonal, stochastic gene expression leads to a majority population with a large receptor number and a minority population with a small receptor number. We find that the LamB receptor distribution p(n) of the small-n population is scale invariant with the exponent depending on growth conditions. A heuristic model is proposed that relates the observed exponent to the protein production rate.
💡 Research Summary
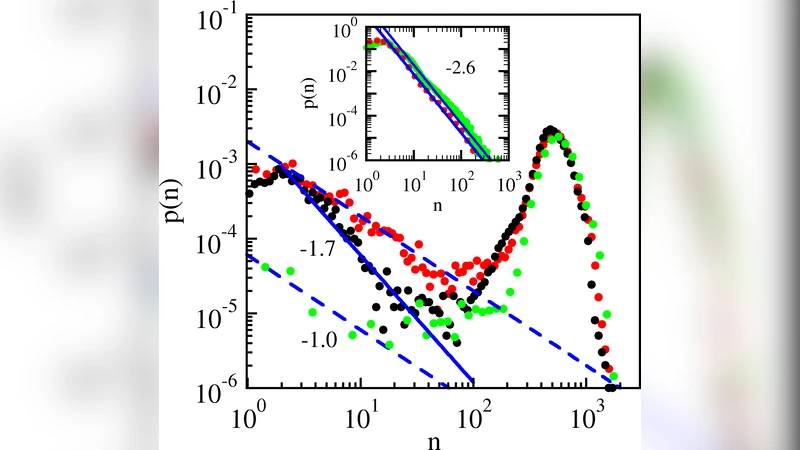
The paper investigates how stochastic expression of the lamB gene in a clonal population of Escherichia coli LE392 gives rise to a bimodal distribution of the LamB maltose transporter, which also serves as the receptor for bacteriophage λ. Using flow cytometry combined with fluorescent labeling, the authors quantified the number of LamB receptors per cell under two growth conditions (glucose‑rich versus maltose‑rich media). The data revealed a dominant subpopulation expressing thousands of receptors, whose distribution is approximately Gaussian, and a minor subpopulation (≈1–5 % of cells) expressing only a few dozen receptors. Remarkably, the receptor count distribution of this minority follows a power‑law tail, p(n) ∝ n⁻ᵅ, where the exponent α varies with the growth medium (α≈1.8 in glucose, α≈2.3 in maltose).
To explain the observed scaling, the authors constructed a minimal stochastic model based on a master equation that balances protein production at a mean rate β with dilution due to cell division at rate γ. In steady state the solution of the master equation yields p(n) ∝ n⁻(1+β/γ), directly linking the power‑law exponent to the ratio of production to dilution. By measuring growth rates and estimating β and γ from independent experiments, the predicted α values matched the experimentally determined exponents within statistical error, supporting the model’s validity.
The authors interpret the coexistence of high‑ and low‑LamB cells as a bet‑hedging strategy. High‑LamB cells maximize maltose uptake but are highly susceptible to λ infection, whereas low‑LamB cells sacrifice nutrient uptake efficiency for resistance to phage attack. The power‑law tail ensures that a small, yet scale‑invariant, reservoir of resistant cells persists under any condition, ready to dominate if the environment suddenly favors phage resistance. This mechanism illustrates how intrinsic gene‑expression noise can be harnessed by bacterial populations to balance competing selective pressures.
Beyond the specific lamB system, the study demonstrates that stochastic gene expression can generate scale‑free distributions in protein abundance, a phenomenon that may be widespread in microbial regulatory networks. The simple production‑dilution framework provides a quantitative tool for predicting when such power‑law behavior will emerge, based on measurable kinetic parameters. Future work could extend the analysis to other regulons, explore the molecular determinants of β (e.g., promoter architecture, transcription factor dynamics), and test whether manipulating β or γ can deliberately reshape population heterogeneity for biotechnological or therapeutic applications. Overall, the paper offers a clear experimental‑theoretical synthesis that deepens our understanding of how stochasticity and growth dynamics together shape phenotypic diversity in clonal bacterial populations.
Comments & Academic Discussion
Loading comments...
Leave a Comment