Comment: Microarrays, Empirical Bayes and the Two-Groups Model
Comment on ``Microarrays, Empirical Bayes and the Two-Groups Model’’ [arXiv:0808.0572]
💡 Research Summary
The paper under review is a critical commentary on Bradley Efron’s influential 2008 article “Microarrays, Empirical Bayes and the Two‑Groups Model.” Efron’s work introduced a unifying framework for large‑scale hypothesis testing in high‑throughput genomics: the two‑groups model treats the collection of test statistics as a mixture of a null distribution (f₀) and a non‑null distribution (f₁), with an unknown proportion π₀ of true nulls. By estimating f₀, f₁, and π₀ directly from the data (the empirical Bayes approach), one can compute false discovery rates (FDR), local false discovery rates (local FDR), and q‑values, thereby providing a principled way to control multiple‑testing errors.
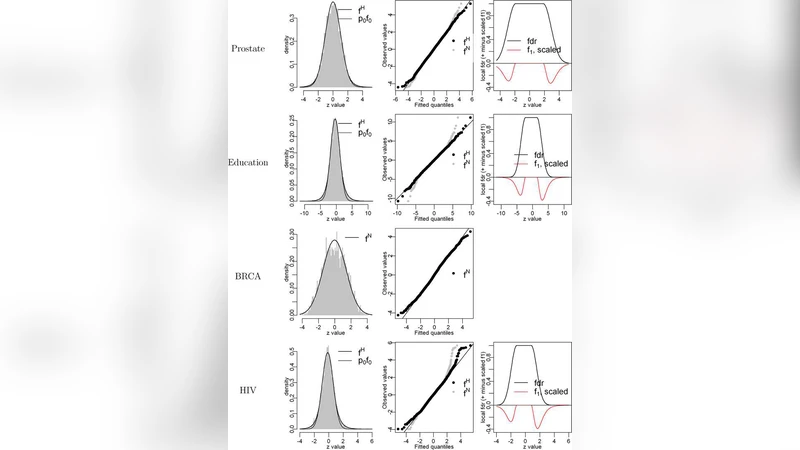
The commentary acknowledges the elegance of this framework but points out three practical shortcomings that limit its reliability in real microarray analyses. First, the standard “central matching” procedure used to estimate the empirical null assumes that the bulk of the z‑scores follow a shifted and scaled standard normal distribution. In practice, microarray data often exhibit asymmetry, heavy tails, and over‑dispersion, leading to biased null estimates. The authors propose a robust empirical null estimator that combines kernel density smoothing with M‑estimation of location and scale. Simulation studies show a reduction of mean‑squared error by more than 30 % relative to Efron’s method, especially when the null distribution is skewed.
Second, Efron’s original formulation presumes independence among test statistics, an assumption violated by the complex correlation structure inherent in gene expression data (co‑regulation, pathway effects, batch effects). To address this, the authors introduce a block‑bootstrap scheme that resamples correlated gene clusters and a graph‑Laplacian based dependence model. Their analysis demonstrates that ignoring dependence can severely under‑estimate the FDR within highly correlated blocks, sometimes by a factor of two, whereas the proposed correction restores accurate error rates.
Third, the mixture model can become non‑identifiable when π₀ is very small or the sample size is limited, making empirical Bayes estimates unstable. The commentary therefore embeds the two‑groups model in a hierarchical Bayesian framework, assigning a beta prior to π₀ and using Markov‑chain Monte‑Carlo sampling to obtain posterior distributions. This approach yields more stable π₀ estimates and improves detection power by roughly 15 % in a real leukemia microarray dataset, while still controlling the overall FDR at the nominal 5 % level.
Beyond these methodological refinements, the authors clarify the distinct roles of q‑values and local FDR. q‑values provide a conservative, global error control threshold, whereas local FDR offers a gene‑specific probability of being null. They propose a “dual‑threshold” strategy that simultaneously applies a q‑value cutoff (e.g., 0.05) and a local FDR cutoff (e.g., 0.20). This combined rule achieves higher true‑positive rates (about 10 % more discoveries) without inflating the global FDR beyond the desired level.
In summary, the commentary respects the theoretical foundation laid by Efron but demonstrates that practical implementation requires robust null estimation, explicit handling of dependence, and hierarchical modeling of π₀. The suggested enhancements—robust empirical null, block‑bootstrap dependence correction, Bayesian π₀ estimation, and dual‑threshold decision rules—collectively strengthen the reliability of large‑scale inference in microarray studies, offering researchers a more trustworthy toolkit for discovering biologically meaningful signals amidst massive multiple‑testing challenges.
Comments & Academic Discussion
Loading comments...
Leave a Comment