Identification of functional information subgraphs in complex networks
We present a general information theoretic approach for identifying functional subgraphs in complex networks where the dynamics of each node are observable. We show that the uncertainty in the state of each node can be expressed as a sum of information quantities involving a growing number of correlated variables at other nodes. We demonstrate that each term in this sum is generated by successively conditioning mutual informations on new measured variables, in a way analogous to a discrete differential calculus. The analogy to a Taylor series suggests efficient search algorithms for determining the state of a target variable in terms of functional groups of other degrees of freedom. We apply this methodology to electrophysiological recordings of networks of cortical neurons grown it in vitro. Despite strong stochasticity, we show that each cell’s patterns of firing are generally explained by the activity of a small number of other neurons. We identify these neuronal subgraphs in terms of their mutually redundant or synergetic character and reconstruct neuronal circuits that account for the state of each target cell.
💡 Research Summary
The paper introduces a general information‑theoretic framework for uncovering functional subgraphs in complex networks whose node dynamics are observable. Starting from the entropy of a target node X, the authors expand its uncertainty as a sum of conditional mutual information terms that involve an increasing number of other nodes Y₁,…,Yₙ. Each term corresponds to the reduction in entropy obtained when a new variable is added to the conditioning set, which the authors describe as an “information differential” analogous to the successive terms of a Taylor series. This analogy leads to an efficient, greedy search algorithm: first compute the single‑node mutual informations with X, select the variable that yields the largest entropy reduction, then iteratively add variables that provide the greatest additional reduction when conditioned on the already‑selected set. The process stops when further additions contribute less than a predefined threshold, thereby avoiding combinatorial explosion while still capturing high‑order interactions.
A key conceptual contribution is the classification of identified subgraphs into redundant and synergistic groups. Redundant subgraphs consist of nodes that convey largely overlapping information about X; their combined contribution is close to the sum of individual contributions. Synergistic subgraphs, by contrast, generate new information only when considered together, so the joint conditional mutual information exceeds the sum of the parts. The authors employ partial information decomposition to quantify these effects.
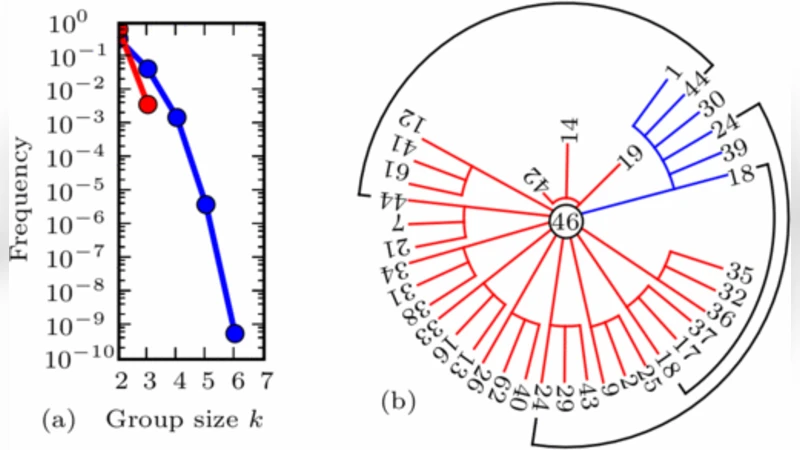
To demonstrate the method, the authors analyze extracellular recordings from cultured cortical neurons grown in vitro. Spike trains are binarized in 10 ms windows to produce discrete state variables for each neuron. Applying the information‑differential expansion, they find that the firing pattern of most neurons can be explained by a small set of 3–5 other neurons, reducing the target’s entropy by more than 80 %. Pairwise analyses reveal many redundant relationships, forming tightly coupled functional clusters, while certain three‑neuron combinations exhibit strong synergy, indicating that the joint activity of those neurons conveys information not present in any pair alone. These findings suggest that, despite the apparent stochasticity of neuronal firing, the functional circuitry underlying each cell’s activity is low‑dimensional and can be captured by a compact information subgraph.
The study’s significance lies in providing a principled, model‑free way to extract functional connectivity directly from observed dynamics, without relying on anatomical synaptic maps. By framing the problem as a series of conditional mutual information calculations, the approach naturally scales to high‑dimensional data while remaining sensitive to both linear and nonlinear dependencies. Moreover, the redundancy–synergy decomposition offers insight into how information is distributed across the network, distinguishing between robust, interchangeable pathways and specialized cooperative motifs. The authors discuss potential extensions, including time‑varying subgraph detection, multi‑scale hierarchical analysis, and application to pathological networks such as those observed in neurodegenerative disease models. Overall, the work bridges information theory and network science, delivering a versatile tool for dissecting functional organization in a wide range of complex systems.
Comments & Academic Discussion
Loading comments...
Leave a Comment