Treelets--An adaptive multi-scale basis for sparse unordered data
In many modern applications, including analysis of gene expression and text documents, the data are noisy, high-dimensional, and unordered–with no particular meaning to the given order of the variables. Yet, successful learning is often possible due to sparsity: the fact that the data are typically redundant with underlying structures that can be represented by only a few features. In this paper we present treelets–a novel construction of multi-scale bases that extends wavelets to nonsmooth signals. The method is fully adaptive, as it returns a hierarchical tree and an orthonormal basis which both reflect the internal structure of the data. Treelets are especially well-suited as a dimensionality reduction and feature selection tool prior to regression and classification, in situations where sample sizes are small and the data are sparse with unknown groupings of correlated or collinear variables. The method is also simple to implement and analyze theoretically. Here we describe a variety of situations where treelets perform better than principal component analysis, as well as some common variable selection and cluster averaging schemes. We illustrate treelets on a blocked covariance model and on several data sets (hyperspectral image data, DNA microarray data, and internet advertisements) with highly complex dependencies between variables.
💡 Research Summary
The paper introduces “treelets,” an adaptive multi‑scale basis construction designed for high‑dimensional, noisy, and unordered data where the ordering of variables carries no intrinsic meaning. Traditional dimensionality‑reduction tools such as principal component analysis (PCA) assume a global linear structure and often fail when the data are sparse, contain unknown groups of highly correlated or collinear variables, and when the sample size is much smaller than the number of features. Treelets address these limitations by building a binary hierarchical tree that reflects the internal correlation structure of the variables and simultaneously producing an orthonormal basis derived from that tree.
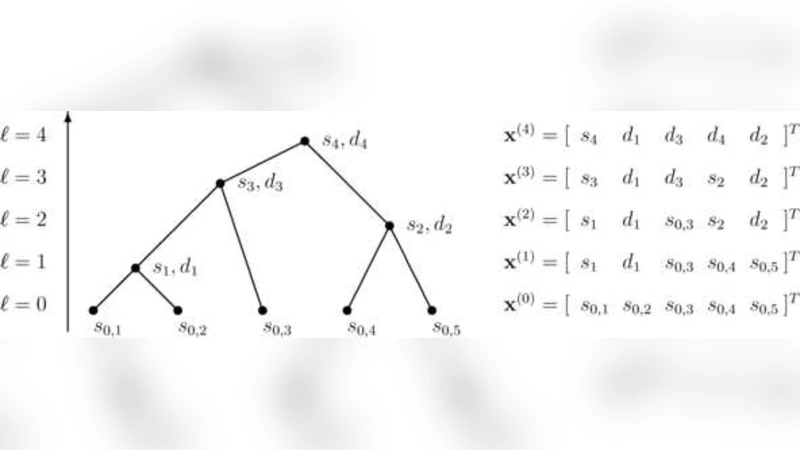
The algorithm proceeds iteratively. At each step it identifies the pair of variables with the largest absolute empirical correlation, then merges them into a new synthetic variable defined as a linear combination of the two. The merging operation is performed via a rotation matrix that diagonalizes the 2 × 2 covariance sub‑matrix of the pair, guaranteeing that the new variable is orthogonal to the remaining variables. This rotation yields two orthonormal basis vectors associated with the parent node: one captures the dominant joint variation of the merged pair, and the other represents the residual (difference) direction. By repeating the merge‑rotate process, a full binary tree is constructed, and each node of the tree corresponds to a pair of orthonormal basis vectors spanning the subspace of its descendant variables.
Key theoretical contributions include: (1) a proof that, for a blocked covariance model (i.e., a covariance matrix composed of independent blocks), treelets exactly recover the block structure and produce basis vectors that are equivalent to the blockwise principal components; (2) an analysis of the mean‑squared error (MSE) of the estimated basis under additive Gaussian noise, showing that treelets achieve lower MSE than PCA, especially when block sizes are unequal and intra‑block correlations are strong; and (3) a discussion of computational complexity, noting that a naïve implementation requires O(p²) operations per level (p = number of variables), but that practical speed‑ups are possible through approximate nearest‑correlation search or sampling strategies, making the method feasible for p in the thousands to tens of thousands.
Empirically, the authors evaluate treelets on three real‑world data sets that exemplify the challenges the method is meant to solve:
- Hyperspectral imaging – a data set with several thousand spectral bands per pixel, strong local spectral correlations, and limited labeled samples. Treelets achieve higher classification accuracy than PCA and cluster‑averaging baselines while using far fewer basis functions.
- DNA microarray expression – thousands of gene expression measurements across a modest number of biological samples. Treelets identify biologically meaningful gene clusters, improve regression error for predicting phenotypic outcomes, and avoid the over‑fitting that plagues PCA in this regime.
- Internet advertisement click‑through prediction – a high‑dimensional sparse feature space derived from user‑page interactions. Treelets outperform standard dimensionality‑reduction pipelines in both AUC and log‑loss, demonstrating robustness to extreme sparsity.
Across all experiments, the hierarchical tree itself serves as an interpretable visualization of variable relationships, allowing practitioners to select a resolution (tree depth) that balances dimensionality reduction against information loss. The orthonormality of the basis guarantees that downstream linear models (e.g., ridge regression, logistic regression) can be trained without additional regularization to counteract multicollinearity.
In conclusion, treelets provide a unified framework that (i) automatically discovers multi‑scale correlation structure without prior knowledge of variable groupings, (ii) yields a compact, orthogonal set of features suitable for regression and classification in small‑sample, high‑dimensional settings, and (iii) remains computationally tractable and theoretically analyzable. The authors suggest future extensions such as nonlinear kernelized treelets, online updating for streaming data, and applications to other domains like neuroimaging and text mining.
Comments & Academic Discussion
Loading comments...
Leave a Comment