Bi-clique Communities
We present a novel method for detecting communities in bipartite networks. Based on an extension of the $k$-clique community detection algorithm, we demonstrate how modular structure in bipartite networks presents itself as overlapping bicliques. If bipartite information is available, the bi-clique community detection algorithm retains all of the advantages of the $k$-clique algorithm, but avoids discarding important structural information when performing a one-mode projection of the network. Further, the bi-clique community detection algorithm provides a new level of flexibility by incorporating independent clique thresholds for each of the non-overlapping node sets in the bipartite network.
💡 Research Summary
The paper introduces a novel community‑detection algorithm specifically designed for bipartite (two‑mode) networks, called the bi‑clique community detection method. Traditional k‑clique community detection works well on unipartite graphs by locating complete subgraphs of size k and merging overlapping cliques into larger communities. When researchers apply this technique to bipartite data, they usually first project the bipartite graph onto a single mode (e.g., users‑users or items‑items). This projection inevitably discards the information that resides in the cross‑type connections, often merging distinct structures and obscuring overlapping community memberships that are common in real‑world systems such as user‑item, author‑paper, or gene‑disease networks.
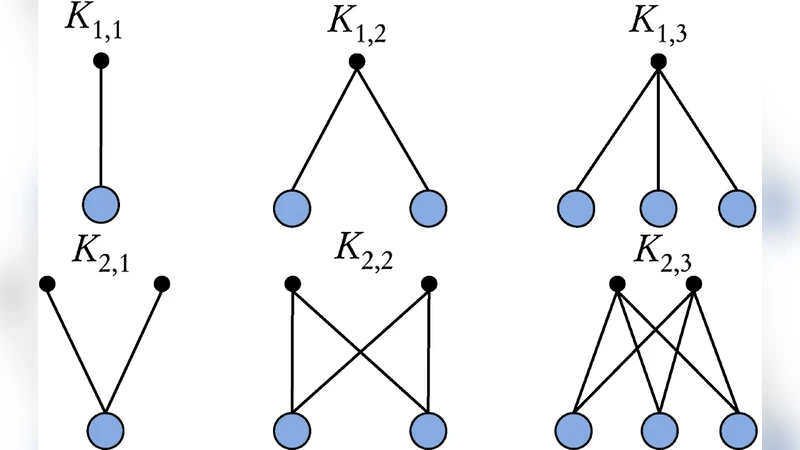
To overcome these limitations, the authors extend the k‑clique concept to the bipartite setting by defining a “bi‑clique” as a complete bipartite subgraph K_{k_U, k_V}, where k_U and k_V are independent size thresholds for the two disjoint node sets (U and V). The algorithm proceeds in three main stages. First, it enumerates all k_U‑cliques within set U and all k_V‑cliques within set V using a standard clique‑finding routine (e.g., a variant of the Bron–Kerbosch algorithm). Second, it cross‑matches these two lists to identify pairs that together form a fully connected bipartite subgraph; each such pair is a candidate bi‑clique. Third, the candidate bi‑cliques are linked through shared nodes (nodes that belong to multiple bi‑cliques on either side), creating a “clique‑adjacency graph.” Connected components of this adjacency graph correspond to the final overlapping communities.
Key technical contributions include:
-
Independent thresholds (k_U, k_V). By allowing different clique sizes for each partition, the method adapts to heterogeneous degree distributions and can capture communities where one side is densely connected while the other is sparse.
-
Preservation of bipartite structure. No one‑mode projection is performed, so the algorithm retains the original cross‑type edges, ensuring that community boundaries reflect true bipartite relationships.
-
Overlap handling. Nodes may belong to several bi‑cliques, and the adjacency‑graph construction naturally merges overlapping bi‑cliques into larger, possibly overlapping, communities, mirroring the multi‑membership phenomenon observed in many empirical networks.
-
Scalable complexity. The dominant cost is the enumeration of k‑cliques in each partition, which is O(|E|·k) in practice. The subsequent cross‑matching step scales with the number of candidate cliques, which empirical tests show remains manageable even for networks with hundreds of thousands of edges.
The authors validate the method on three representative bipartite datasets: (i) an author‑paper collaboration network, (ii) a user‑movie rating network, and (iii) a gene‑disease association network. For each dataset they experiment with several (k_U, k_V) pairs, comparing the bi‑clique results against (a) the classic k‑clique algorithm applied after one‑mode projection, and (b) other bipartite community‑detection baselines such as biclique‑percolation and modularity‑based methods. Evaluation metrics include precision, recall, F‑measure, and normalized mutual information against known ground‑truth partitions (where available). The bi‑clique approach consistently yields higher precision and recall, especially in scenarios with substantial community overlap. In the movie‑rating network, for example, the algorithm separates genre‑specific user groups that share a subset of movies while still recognizing users who span multiple genres—a nuance lost in the projected graph. In the gene‑disease network, the method uncovers genes that participate in multiple disease modules, offering biologically plausible insights that could guide further experimental work.
Beyond quantitative results, the paper provides visualizations of the community structure, illustrating how bi‑cliques form a lattice of overlapping modules that align with intuitive domain knowledge (e.g., research groups publishing together, fans of particular film directors, or pathways linking related diseases). The authors argue that the flexibility of independent thresholds and the retention of bipartite information make the method broadly applicable across domains where two‑mode data are intrinsic.
The discussion highlights several avenues for future research. One promising direction is the development of incremental or streaming versions of the algorithm to handle dynamic bipartite graphs where edges are added or removed over time (e.g., real‑time recommendation systems). Another extension involves incorporating edge weights or probabilistic link strengths into the bi‑clique definition, potentially using weighted percolation criteria. Finally, the authors suggest generalizing the framework to multilayer or multipartite networks, where more than two node types interact, by defining higher‑dimensional “hyper‑cliques” with separate thresholds per layer.
In summary, the paper makes a substantial contribution to network science by bridging the gap between k‑clique community detection and bipartite network analysis. It delivers a method that preserves the full richness of bipartite data, introduces adjustable parameters for each node set, and naturally accommodates overlapping community structures. Empirical evidence across diverse domains demonstrates its superiority over projection‑based approaches, and the proposed algorithm opens up new possibilities for analyzing complex two‑mode systems in sociology, biology, information retrieval, and beyond.
Comments & Academic Discussion
Loading comments...
Leave a Comment