Inferring Neuronal Network Connectivity from Spike Data: A Temporal Datamining Approach
Understanding the functioning of a neural system in terms of its underlying circuitry is an important problem in neuroscience. Recent developments in electrophysiology and imaging allow one to simultaneously record activities of hundreds of neurons. Inferring the underlying neuronal connectivity patterns from such multi-neuronal spike train data streams is a challenging statistical and computational problem. This task involves finding significant temporal patterns from vast amounts of symbolic time series data. In this paper we show that the frequent episode mining methods from the field of temporal data mining can be very useful in this context. In the frequent episode discovery framework, the data is viewed as a sequence of events, each of which is characterized by an event type and its time of occurrence and episodes are certain types of temporal patterns in such data. Here we show that, using the set of discovered frequent episodes from multi-neuronal data, one can infer different types of connectivity patterns in the neural system that generated it. For this purpose, we introduce the notion of mining for frequent episodes under certain temporal constraints; the structure of these temporal constraints is motivated by the application. We present algorithms for discovering serial and parallel episodes under these temporal constraints. Through extensive simulation studies we demonstrate that these methods are useful for unearthing patterns of neuronal network connectivity.
💡 Research Summary
The paper tackles the challenging problem of inferring neuronal connectivity from multi‑neuronal spike‑train recordings, which have become increasingly abundant thanks to advances in micro‑electrode arrays, optical imaging, and related technologies. The authors propose to view a spike train as a sequence of discrete events, each event being a pair (neuron ID, timestamp). Within the framework of temporal data mining, they define “episodes” – patterns of events that occur frequently – and distinguish two principal classes: serial episodes (ordered sequences) and parallel episodes (sets of events occurring within a short time window).
A key innovation is the incorporation of biologically motivated temporal constraints into the episode definition. Constraints such as minimum and maximum inter‑event intervals, overall episode duration, and refractory periods reflect synaptic transmission delays and neuronal recovery times. By imposing these constraints, the method filters out spurious coincidences that would otherwise dominate a naïve frequency count.
To discover constrained frequent episodes efficiently, the authors extend classic Apriori‑style mining algorithms with a window‑based counting mechanism and priority‑queue data structures. For serial episodes, each candidate extension is accepted only if the time gap between the newly added event and its predecessor lies within the prescribed bounds. For parallel episodes, all constituent events must fall inside a predefined sliding window. This pruning dramatically reduces the combinatorial explosion, enabling the analysis of data sets containing hundreds of neurons and tens of thousands of spikes.
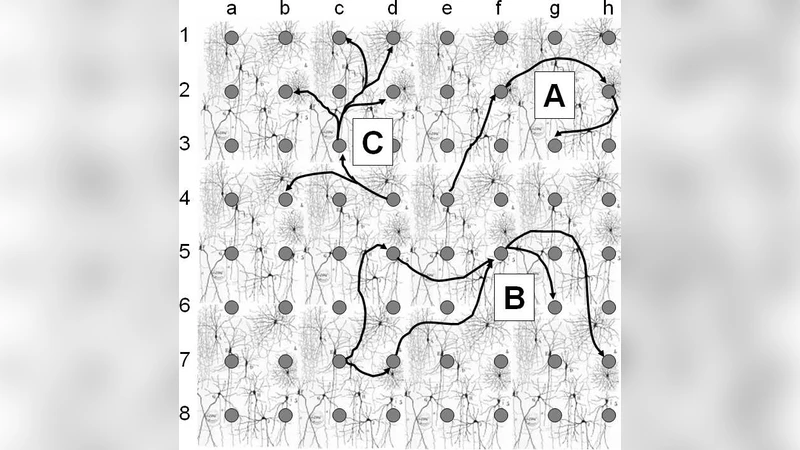
The methodological contribution is validated through extensive simulations. Each neuron is modeled as an inhomogeneous Poisson process whose firing rate is modulated by incoming spikes from presynaptic partners. The simulator allows the user to embed various connectivity motifs – feed‑forward chains, feedback loops, convergent/divergent patterns – and to assign realistic synaptic delays and efficacies. When the mining algorithms are applied to the simulated spike streams, the embedded motifs are recovered as high‑frequency serial or parallel episodes, even when the motifs involve five to ten neurons.
Beyond synthetic data, the authors apply their approach to real spike‑train recordings from cultured cortical networks. The analysis uncovers known synchronous clusters and reveals putative synfire chains, demonstrating that the technique can extract meaningful connectivity information from noisy biological recordings. Statistical significance is assessed by comparing episode frequencies against surrogate data generated through temporal shuffling, confirming that the discovered patterns are unlikely to arise by chance.
The paper highlights several advantages of the proposed framework: (1) model independence – it does not rely on a specific stochastic model of neuronal interaction; (2) scalability – the pruning strategy keeps computational cost near‑linear in the number of events; (3) ease of statistical validation – frequent episodes can be directly tested against null models. Limitations are also acknowledged: fixed temporal windows may miss variable synaptic delays; the Poisson‑based simulator does not capture all biophysical aspects of neuronal firing; and episode frequency alone cannot establish causality, suggesting a need for downstream causal inference methods.
In summary, the authors successfully bridge temporal data mining and neuroinformatics, offering a powerful, efficient, and extensible tool for uncovering high‑order temporal patterns that reflect underlying neuronal circuitry. This work opens avenues for real‑time connectivity monitoring, integration with Bayesian network inference, and potential applications in brain‑machine interfaces and neurological disorder diagnostics.
Comments & Academic Discussion
Loading comments...
Leave a Comment